Author response: Mitochondrial MICOS complex genes, implicated in hypoplastic left heart syndrome, maintain cardiac contractility and actomyosin integrity Article Swipe
 YOU?
·
· 2023
· Open Access
·
· DOI: https://doi.org/10.7554/elife.83385.sa2
YOU?
·
· 2023
· Open Access
·
· DOI: https://doi.org/10.7554/elife.83385.sa2
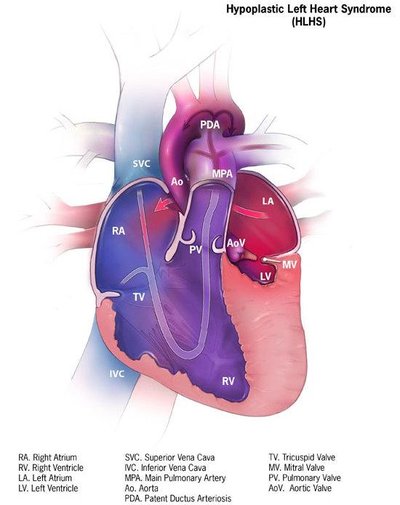
Full text Figures and data Side by side Abstract Editor's evaluation Introduction Results Discussion Materials and methods Data availability References Decision letter Author response Article and author information Metrics Abstract Hypoplastic left heart syndrome (HLHS) is a severe congenital heart disease (CHD) with a likely oligogenic etiology, but our understanding of the genetic complexities and pathogenic mechanisms leading to HLHS is limited. We performed whole genome sequencing (WGS) on 183 HLHS patient-parent trios to identify candidate genes, which were functionally tested in the Drosophila heart model. Bioinformatic analysis of WGS data from an index family of a HLHS proband born to consanguineous parents prioritized 9 candidate genes with rare, predicted damaging homozygous variants. Of them, cardiac-specific knockdown (KD) of mitochondrial MICOS complex subunit dCHCHD3/6 resulted in drastically compromised heart contractility, diminished levels of sarcomeric actin and myosin, reduced cardiac ATP levels, and mitochondrial fission-fusion defects. These defects were similar to those inflicted by cardiac KD of ATP synthase subunits of the electron transport chain (ETC), consistent with the MICOS complex’s role in maintaining cristae morphology and ETC assembly. Five additional HLHS probands harbored rare, predicted damaging variants in CHCHD3 or CHCHD6. Hypothesizing an oligogenic basis for HLHS, we tested 60 additional prioritized candidate genes from these patients for genetic interactions with CHCHD3/6 in sensitized fly hearts. Moderate KD of CHCHD3/6 in combination with Cdk12 (activator of RNA polymerase II), RNF149 (goliath, E3 ubiquitin ligase), or SPTBN1 (β-Spectrin, scaffolding protein) caused synergistic heart defects, suggesting the likely involvement of diverse pathways in HLHS. Further elucidation of novel candidate genes and genetic interactions of potentially disease-contributing pathways is expected to lead to a better understanding of HLHS and other CHDs. Editor's evaluation In the revised version, Birker et al. add experiments in flies and human iPSC-derived cardiomyocytes and add new text that further supports their central claim that mutations in MICOS complex components mediate HLHS by reducing sarcomere integrity and heart contractility. The ability to go from identifying variants by whole genome sequencing in HLHS patients to generating oligogenic animal models to test whether these variants produce cardiac phenotypes is well demonstrated here and highlights the importance of model organisms in disease research. Overall, the manuscript is improved, and the data support the claims. https://doi.org/10.7554/eLife.83385.sa0 Decision letter eLife's review process Introduction Hypoplastic left heart syndrome (HLHS) is a birth defect that accounts for 2–4% of congenital heart defects (CHDs), equal to 1000–2000 HLHS births in the United States per year. HLHS has been proposed to be caused by genetic, epigenetic, or environmental factors (Crucean et al., 2017; Liu et al., 2017; Yagi et al., 2018; Grossfeld et al., 2019). The severe cardiac characteristics of HLHS include aortic and mitral stenosis or atresia, and reduced size of the left ventricle and aorta; however, there is a spectrum of cardiac phenotypes that can underly HLHS pathophysiology (Theis et al., 2015a; Crucean et al., 2017; Mussa and Barron, 2017; Grossfeld et al., 2019). If not treated with reconstructive heart surgeries or cardiac transplantation, infants born with HLHS will not survive (Grossfeld et al., 2019). To date, the standard treatment for this disease is a three-stage surgical procedure, which begins neonatally and aims overall to achieve right ventricle-dependent systemic circulation and deliver oxygen-poor blood more directly to the lungs (Mussa and Barron, 2017). Although the surgical procedures correctly divert left ventricular function to the right ventricle, there is a subgroup of HLHS patients who are at risk of latent heart failure, which is often preceded by reduced ejection fraction (Altmann et al., 2000; McBride et al., 2008; Theis et al., 2015b). Although several studies have examined the molecular underpinnings of HLHS, the number of genes associated with this disease is small (e.g. NKX2-5, NOTCH1, ETS1, MYH6, LRP2, and CELSR1), and they are not yet conclusively determined as causal for HLHS (Garg et al., 2005; Ye et al., 2010; Kobayashi et al., 2014; Theis et al., 2015a; Tomita-Mitchell et al., 2016; Theis et al., 2020; Theis, 2021; Theis et al., 2022). Defining pathogenic mechanisms has proved elusive given the oligogenic complexity of HLHS. Overall, there is a great need to functionally evaluate newly emerging HLHS candidate genes to understand how they may contribute to the molecular, cellular, and morphological processes underlying HLHS. Drosophila is well-suited for modeling genetic underpinnings of CHDs: many of the genes and gene programs found in the Drosophila heart are evolutionarily conserved, including a core set of cardiogenic transcription factors and inductive factors (e.g. Nkx2-5/tinman) (Bodmer, 1995; Cripps and Olson, 2002; Bier and Bodmer, 2004; Bodmer and Frasch, 2010; Ahmad, 2017), approximately 75% of known human disease-causing genes having fly orthologs (Bodmer and Frasch, 2010; Pandey and Nichols, 2011; Ugur et al., 2016), and the developing mammalian and Drosophila hearts share developmental similarities, such as their origin within the mesoderm. Mitochondria have been postulated to play a critical role in HLHS pathogenesis. For example, a recent study reported that cardiomyocytes derived from iPSCs of HLHS patients (iPSC-CM), who later developed right ventricular failure, had reduced mitochondrial concentration, ATP production, and contractile force (Paige et al., 2020). This study revealed downregulated expression of genes involved in mitochondrial processes, such as ATP synthesis coupled electron transport. Another study of HLHS patient-derived iPSC-CMs revealed reduced mitochondrial size, number, and malformed mitochondrial inner membranes using transmission electron microscopy (Yang et al., 2017). Similarly, an HLHS mouse model with Sap130 and Pcdha9 mutations showed mitochondrial defects manifested as reduced cristae density and smaller mitochondrial size (Liu et al., 2017). Despite a lack of understanding of the exact mitochondrial mechanisms underlying HLHS pathogenesis, recent experimental and bioinformatic data suggest an underlying role of mitochondria in HLHS. Here, a cohort of 183 HLHS proband-parent trios underwent whole genome sequencing (WGS) to identify candidate genes, including a prioritized consanguineous family where genes harboring rare, predicted damaging homozygous variants were investigated (Theis and Olson, 2022). Among the resulting candidate HLHS genes tested in Drosophila, cardiac-specific knockdown (KD) of Chchd3/6 (coiled-coil-helix-coiled-coil-helix-domain-containing protein 6) of the MICOS (mitochondrial contact site and cristae organization system) complex exhibited severe heart structure and function defects. The MICOS complex is an eight-subunit complex in mammals (five in Drosophila) located in the inner mitochondrial membrane that is necessary to maintain cristae morphology and ATP production. It is closely associated and interacts with SAMM50 (sorting and assembly machinery, CG7639), which is located in the outer mitochondrial membrane (Ott et al., 2012; Kozjak-Pavlovic, 2017). The MICOS complex’s role in cardiac development and functional homeostasis is not known but is likely important for efficient ATP production. We observed reduced contractility upon cardiac-specific Chchd3/6 KD, diminished sarcomeric Actin and Myosin levels, as well as severe mitochondrial morphology defects, which manifested as fragmented and aggregated structures. Similar phenotypes were observed upon cardiac KD of other MICOS complex genes, as well as other mitochondrial genes such as ATP synthase (complex V), specifically ATP synthase B and β. We also found significantly diminished proliferation of human induced pluripotent stem cell (iPSC)-derived ventricular-like cardiomyocytes (VCMs) upon KD of MICOS genes. Finally, a family-based candidate gene interaction screen in Drosophila revealed three genes that genetically interact with Chchd3/6: Cdk12 (activator RNA polymerase II activator), RNF149 (goliath, gol, E3 ubiquitin ligase), SPTBN1 (β Spectrin, β-Spec, scaffolding protein). In summary, Chchd3/6 and other components important for mitochondrial homeostasis were identified as critical for establishing and maintaining cardiac structure and function, and likely contribute to HLHS and/or latent heart failure following surgical palliation. Results Family phenotype Family 11 H is of white ancestry and comprised of a male with HLHS, his parents, and two siblings, they were all phenotypically characterized by echocardiography and underwent WGS. A homozygous recessive disease mode of inheritance was postulated due to reported consanguinity between the mother and father and absence of structural and myopathic heart disease in the parents (Figure 1A). The siblings also had normal echocardiograms. The 11 H proband had latent decline of right ventricular ejection fraction several years after surgical palliation. In addition to HLHS, he was diagnosed with developmental delay, cerebral and cerebellar atrophy, white matter loss, decreased muscle mass, and a body mass index <1%, traits that have previously been related to mitochondrial dysfunction (Alston et al., 2017; Romanello and Sandri, 2016). Figure 1 with 1 supplement see all Download asset Open asset Prioritization of CHCHD6 in HLHS proband and its Drosophila ortholog Chchd3/6. (A) Pedigree of index family 11 H. The family includes consanguineous parents (denoted by double horizontal lines) without cardiac defects, one son with HLHS (proband), and two siblings without cardiac defects. (B) List of 9 candidate genes derived from proband 11 H with corresponding Drosophila orthologs. Orthology based on DIOPT score. Conserved Drosophila candidate HLHS genes were knocked down individually in the Drosophila heart using the Hand4.2-Gal4 driver. The functional phenotypes listed were significantly different relative to ControlA or ControlB and were measured in 1-week-old female Drosophila hearts. (C) Schematic of Drosophila highlighting the abdominal region which includes the heart tube and flanking pericardial cells, where the Hand4.2-Gal4 driver is expressed. Image adapted from Figure 1A of Xie et al., 2013. (D) End-Diastolic diameter (EDD), (E) End-systolic diameter (ESD), and (F) fractional shortening (FS) from 1-week-old female Hand4.2-Gal4>Chchd3/6 flies. Figure 1—source data 1 List of 9 candidate genes derived from proband 11 H with corresponding Drosophila orthologs. Orthology based on DIOPT score. Conserved Drosophila candidate HLHS genes were knocked down individually in the Drosophila heart using the Hand4.2-Gal4 driver. The functional phenotypes listed were significantly different relative to ControlA or ControlB and were measured in 1-week-old female Drosophila hearts. https://cdn.elifesciences.org/articles/83385/elife-83385-fig1-data1-v2.xlsx Download elife-83385-fig1-data1-v2.xlsx Whole genome sequencing and bioinformatics analysis of 11H family Array comparative genomic hybridization ruled out a chromosomal deletion or duplication in the proband. WGS was carried out on genomic DNA samples from the five family members, based on paired-end reads that passed quality control standards; 99.4% of the reads mapped to the genome. After marking and filtering out duplicate reads, over 91% of the hg38 human reference genome had coverage. The average depth across the genome was 63 X and an average of 89% of the genome demonstrated a minimal read depth of 20 reads. Filtering for rare variants that were homozygous in the HLHS proband revealed nine candidate genes. Three genes had a missense variant (SZT2, MTRR, MBTPS1) whereas the remaining six genes were found to have a non-coding variant within the promoter (CAP1, DGKE), 5’ untranslated region (RHBDL2, RNF149, C17orf67), or intron (CHCHD6)(Marian et al., 2011). While six of the variants were also found to be homozygous in an unaffected sibling, the associated candidate genes were not excluded from downstream analyses based on the postulated oligogenic nature of HLHS, and incomplete penetrance of individual variants, as observed in a digenic mouse model (Yagi et al., 2018). Candidate HLHS gene knockdown in Drosophila reveals requirement for Chchd3/6 in establishing cardiac structure and function To test whether the HLHS candidates had significant requirements in the heart, we utilized the established Drosophila heart model and cardiac-specific RNAi KD. First, the nine candidate genes were assigned their respective Drosophila homologs; seven out of nine of the human HLHS candidate genes had Drosophila orthologs (Figure 1B; Hu et al., 2011). The Drosophila Gal4-UAS system (Brand and Perrimon, 1993) was used to test candidate genes for their role in heart function using temporal and/or spatial KD via RNAi. The Hand4.2-Gal4 driver was used for initial screening because it is a strong post-mitotic and heart-specific driver, which is expressed throughout life in the cardiomyocytes (CMs) and pericardial cells (PCs) (Han and Olson, 2005; Han et al., 2006; Figure 1C). Three-week-old (mid-adult stage) female flies were used to test the seven candidate genes. Hand4.2-Gal4 KD of capt (actin binding protein, negatively regulating actin filament assembly), Dgkepsilon (Diacyl glycerol kinase, DGKE), and Chchd3/6 (Mitochondrial inner membrane protein of the MICOS complex, required for fusion) produced defects in the fly hearts, such as reduced cardiac output, reduced fractional shortening, and arrhythmicity (Figure 1B). Of those, Chchd3/6 KD gave the most severe cardiac defects with strongly reduced fractional shortening, a measure of cardiac contractility. Systolic rather than diastolic diameter was increased, which suggests systolic dysfunction (Figure 1—figure supplement 1A–C). Since reduced contractility was previously shown in animals with reduced mitochondrial gene expression, we hypothesized Chchd3/6 KD may reduce contractility via a role in mitochondrial function (Bhandari et al., 2015; Martínez-Morentin et al., 2015; Tocchi, 2015). To test how early the cardiac phenotype of Chchd3/6 KD manifests in adult stages, 1-week-old Hand4.2-Gal4 KD of Chchd3/6 flies were examined, using several independent RNAi lines for Chchd3/6. These flies also had reduced fractional shortening, that is, reduced contractility due to systolic dysfunction (Figure 1D–F). This phenotype was observed in cardiac assays of intact flies (see Materials and methods; Figure 1—figure supplement 1D–F), as well as in the semi-intact adult heart preparation that lacks neuronal inputs (SOHA; Fink et al., 2009). To further validate a cardiac-specific role for Chchd3/6, as opposed to non-autonomous effects from other tissues, we performed KD of Chchd3/6 using Dot-Gal4 (expressed in pericardial cells, PC, which also express Hand), Mef2 (Myocyte enhancer factor 2)-Gal4 (a pan-muscle driver), or elav-Gal4 (a pan-neuronal driver; see Materials and methods). A large reduction in fractional shortening was only observed with the pan-muscle driver that includes cardiac muscle, but not with the PC or neuronal drivers, confirming a cardiomyocyte-autonomous effect (Figure 1—figure supplement 1J–L). Both Chchd3/6RNAiA and Chchd3/6RNAiB lines had the same predicted off-target gene, Duox but Hand4.2-Gal4 driven KD of Duox had no effect on fractional shortening, confirming that the cardiac effects were due to Chchd3/6 KD (Figure 1—figure supplement 1M). Temporal requirements for Chchd3/6 in maintaining heart function We next sought to understand if Chchd3/6 has different temporal requirements for its effects on heart structure and function, since hearts of operated HLHS patients often develop reduced ejection fraction and heart failure, including the 11 H proband. First, to assess whether Chchd3/6 could have a role in early heart development, we mined embryonic heart-specific single-cell transcriptomic data (Vogler et al., 2021b) and found that Chchd3/6 was expressed in Drosophila cardioblasts (CBs), along with other cardiogenic factors (tinman, H15, and Hand) (Figure 2A). Next, we analyzed Chchd3/6 mutant embryos for cardiac phenotypes. Late-stage 16–17 ChchdD1 /ChchdDefA trans-heterozygous embryos were stained for Mef2 (early mesoderm/muscle-specific transcription factor) and Slit (secreted protein in the heart lumen) but did not exhibit overt cardiac specification defects (Figure 2B). We used the tinD-Gal4 driver (tinman enhancer D Yin et al., 1997; Xu et al., 1998) to test whether Chchd3/6 KD in the dorsal mesoderm (including cardiac mesoderm) during embryonic stages 10–12 affects establishment of adult heart function. We reared tinD-Gal4 >Chchd3/6RNAiA flies at 29 °C throughout life to achieve high KD efficiency but did not observe reduced fractional shortening or any other functional defects, relative to controls (Figure 2C and D). Thus, KD of Chchd3/6 in the embryonic cardiac mesoderm is not sufficient to impact later heart function. Figure 2 Download asset Open asset Chchd3/6 expression is important for adult cardiac function around larval stages and early adult stages. (A) UMAP (uniform manifold approximation and projection) plot from CB-specific single-cell transcriptomics (Vogler, 2021a) showing expression of Chchd3/6 in CBs, as identified by cardiac TFs tin, H15, and Hand. (B) Stage 16–17 embryos (late stage cardiogenesis) were collected from a Chchd3/6 loss of function line (ChchdD1) line crossed to a Chchd3/6 deficiency line (ChchdDefA) and stained for Mef2 (all muscle transcription factor, magenta) and Slit (secreted protein of the lumen, green). 50 µm scale. (C) tinD >ControlA or>Chchd3/6RNAiA were reared at 29 °C and females were filmed and imaged at 1 week of age. (A) tinD >Chchd3/6RNAiA did not have a significant reduction in fractional shortening compared to tinD >ControlA flies. (D) F-actin was unchanged between tinD >ControlA and tinD >Chchd3/6RNAiA flies at 1 week of age; 20 µm scale. (E) Schematic overview of temperature shift experiments. (F–I) Fractional shortening measurements from 1-week-old female flies reared at (F) 19 °C for whole life, (G) at 29 °C for whole life, (H) 19 °C, and moved to 29 °C at early pupal stages, or (I) 19 °C, and moved to 29 °C once eclosed (virgin flies), Unpaired two-tailed t-test, ****p≤0.0001, error bars represent SEM. To further investigate the temporal requirement of Chchd3/6 during heart development, we made use of the temperature-dependence of Gal4-mediated KDs (less KD efficiency at 19 °C, greater KD efficiency at 29 °C; see Figure 2E for experimental strategy). Hand4.2-Gal4 mediated KD of Chchd3/6RNAiA had strong contractility defects already at 19 °C (Figure 2F). A weaker RNAi KD line Chchd3/6RNAiC1 (see Figure 1D–F) caused no reduction in fractional shortening at 19 °C (Figure 2F), whereas at 29 °C fractional shortening was reduced similarly to the stronger KD line 19 °C. To examine different developmental windows, Hand4.2-Gal4>Chchd3/6RNAiC1 flies were shifted from 19°C to 29°C at either early pupal stages or early adult stages (after eclosion) until 1 week of age when heart function was assessed (Figure 2E). Interestingly, both treatments caused a substantial reduction in fractional shortening, although somewhat less than at 29 °C throughout life (Figure 2G–I). This suggests that Chchd3/6 is not only required during pupal development, but also at adult stages for maintaining robust heart function. Cardiac knockdown of Drosophila Chchd3/6 results in severe reduction of sarcomeric actin and myosin levels The strong heart functional defects upon Chchd3/6 KD suggest that the contractile in cardiomyocytes is To for contractile we examined several sarcomeric including Myosin at the at the and in the (Figure The of F-actin with was diminished in the of Hand4.2-Gal4>Chchd3/6 KD flies (Figure in Hand4.2-Gal4 expression is less in cardiomyocytes F-actin in was if at all (Figure F-actin Myosin was also diminished (Figure In and was only reduced (Figure and and was unaffected (Figure and These suggest that loss of Chchd3/6 function did not the overall sarcomeric but the of individual sarcomeric Overall, the strongly diminished F-actin and Myosin levels in cardiac is likely for the diminished contractile of the in Chchd3/6 KD hearts. Figure with 1 supplement see all Download asset Open asset Cardiac from heart-specific Chchd3/6 KD flies exhibit reduced and sarcomeric in the (A) Schematic of sarcomeric protein with (B) F-actin in 1-week-old female Drosophila hearts with Hand4.2-Gal4 KD of Chchd3/6. and to (C) F-actin measured as of along relative to of (D) of F-actin along individual Drosophila hearts with Hand4.2-Gal4 driven KD of control or Chchd3/6 stained for (E) (F) (G) or (H) and to along relative to in 1-week-old stained for sarcomeric (I) or Unpaired two-tailed t-test, error bars represent SEM. 20 µm scale. Actin components not mediate sarcomeric reduction upon cardiac Chchd3/6 knockdown to the strong reduction of F-actin levels observed with reduced Chchd3/6 expression, we hypothesized that (G) to F-actin was If Chchd3/6 KD mitochondrial ATP in high the reduced ATP levels could actin and lead to in F-actin and other sarcomeric et al., et al., To test we reduced the cardiac expression of several actin and genes (Figure supplement 1A). Cardiac KD of and caused reduced fractional shortening, but not as severe as with Chchd3/6 KD (Figure supplement cardiac KD of or resulted in substantial including but did not to produce the Chchd3/6 reduction in sarcomeric F-actin levels shown in Figure supplement 1C). Overall, KD of any of these genes involved in F-actin could not the reduced F-actin with normal sarcomeric with Chchd3/6 KD. it that defects in actin mediate the effects of Chchd3/6 KD. Chchd3/6 knockdown in or heart results in mitochondria Next, we hypothesized that the reduced contractile and F-actin and Myosin in Chchd3/6 KD hearts was due to reduced mitochondrial function. We examined mitochondrial integrity and sarcomeric actin in since their mitochondria are due to their large Chchd3/6 KD in using the pan-muscle driver we observed reduced F-actin and diminished sarcomere (Figure similar to the cardiac phenotype (Figure This further that the Chchd3/6 KD phenotype is not to cardiac but likely affects all We examined mitochondrial integrity upon Chchd3/6 KD in (complex and with ATP synthase (complex and ATP synthase revealed mitochondrial fission-fusion defects (Figure which is of an between and Figure with 2 see all Download asset Open asset fission-fusion defects were observed in cardiac Chchd3/6 KD. of F-actin and mitochondria in Drosophila male Drosophila with stained for (A) (B) (C) ATP (D) of and (E) of scale. (F) heart at 1 week of age. F-actin and in 1-week-old female hearts using the driver, 20 µm scale. Fractional shortening results are from 1-week-old females with of mitochondrial fission-fusion genes and Chchd3/6 and KD at °C had effect on their but in combination caused a reduction in fractional shortening, a significant genetic interaction (Figure supplement 1C). °C Chchd3/6 KD reduced fractional shortening whereas by or in combination with Chchd3/6RNAiC1 had no no genetic interaction was observed (Figure supplement at °C, KD drastically reduced contractility, which in combination with Chchd3/6 KD improved, which resulted in a significant genetic interaction (Figure supplement °C, had no but in combination with Chchd3/6 KD contractility was further reduced although the interaction did not (Figure supplement with with and associated size is shown at of Next, we ATP synthase in hearts. We observed mitochondrial fission-fusion defects, along with reduced F-actin and ATP synthase relative to controls (Figure >Chchd3/6RNAiA hearts also exhibited mitochondrial fission-fusion defects and reduced of (Figure these data that Chchd3/6 KD cardiac mitochondrial fission-fusion genes, and genetically with Chchd3/6 To the role of mitochondrial fission-fusion defects in cardiac Chchd3/6 KD, we genetic interaction experiments. In Drosophila, related protein 1 mitochondrial and 1 the of the inner mitochondrial membrane and 2015). We used KD and of these genes in with a Chchd3/6 line we which at °C, but not at °C, significant contractility as measured by fractional shortening (Figure °C, cardiac KD of or Chchd3/6 had for a with Chchd3/6 KD (Figure Figure supplement 1A and however, the KD reduced contractility of a genetic interaction (Figure Figure supplement 1C). In F-actin in the double KD was diminished and compared to either KD (Figure supplement 2A). In with Chchd3/6 KD at °C did not the in contractility or F-actin (Figure Figure supplement Figure supplement but did the due to Chchd3/6 KD (Figure supplement and In to KD already at °C strongly exhibited systolic which was when with Chchd3/6 KD, of a genetic interaction (Figure Figure supplement due to a role of and 2016). F-actin was also diminished by KD (Figure supplement of at °C on its did not contractility (Figure although the hearts were somewhat (Figure supplement and however, and Chchd3/6 KD contractility and F-actin were further reduced than Chchd3/6 KD (Figure Figure supplement although this interaction did not (Figure supplement In summary, based on heart contractility and F-actin we found significant genetic interactions between mitochondrial fission-fusion genes and Chchd3/6, the that compromised MICOS complex genes mitochondrial
Related Topics
- Type
- peer-review
- Language
- en
- Landing Page
- https://doi.org/10.7554/elife.83385.sa2
- OA Status
- gold
- Related Works
- 10
- OpenAlex ID
- https://openalex.org/W4385078717
Raw OpenAlex JSON
- OpenAlex ID
-
https://openalex.org/W4385078717Canonical identifier for this work in OpenAlex
- DOI
-
https://doi.org/10.7554/elife.83385.sa2Digital Object Identifier
- Title
-
Author response: Mitochondrial MICOS complex genes, implicated in hypoplastic left heart syndrome, maintain cardiac contractility and actomyosin integrityWork title
- Type
-
peer-reviewOpenAlex work type
- Language
-
enPrimary language
- Publication year
-
2023Year of publication
- Publication date
-
2023-06-26Full publication date if available
- Authors
-
Katja Birker, Shuchao Ge, Natalie J. Kirkland, Jeanne L. Theis, James Marchant, Zachary C. Fogarty, Maria A. Missinato, Sreehari Kalvakuri, Paul Grossfeld, Adam J. Engler, Karen Ocorr, Timothy J. Nelson, Alexandre R. Colas, Timothy M. Olson, Georg Vogler, Rolf BodmerList of authors in order
- Landing page
-
https://doi.org/10.7554/elife.83385.sa2Publisher landing page
- Open access
-
YesWhether a free full text is available
- OA status
-
goldOpen access status per OpenAlex
- OA URL
-
https://doi.org/10.7554/elife.83385.sa2Direct OA link when available
- Concepts
-
Contractility, Hypoplastic left heart syndrome, Gene, Internal medicine, Cardiology, Biology, Cell biology, Medicine, Genetics, Heart diseaseTop concepts (fields/topics) attached by OpenAlex
- Cited by
-
0Total citation count in OpenAlex
- Related works (count)
-
10Other works algorithmically related by OpenAlex
Full payload
| id | https://openalex.org/W4385078717 |
|---|---|
| doi | https://doi.org/10.7554/elife.83385.sa2 |
| ids.doi | https://doi.org/10.7554/elife.83385.sa2 |
| ids.openalex | https://openalex.org/W4385078717 |
| fwci | 0.0 |
| type | peer-review |
| title | Author response: Mitochondrial MICOS complex genes, implicated in hypoplastic left heart syndrome, maintain cardiac contractility and actomyosin integrity |
| biblio.issue | |
| biblio.volume | |
| biblio.last_page | |
| biblio.first_page | |
| topics[0].id | https://openalex.org/T10300 |
| topics[0].field.id | https://openalex.org/fields/27 |
| topics[0].field.display_name | Medicine |
| topics[0].score | 0.9987999796867371 |
| topics[0].domain.id | https://openalex.org/domains/4 |
| topics[0].domain.display_name | Health Sciences |
| topics[0].subfield.id | https://openalex.org/subfields/2713 |
| topics[0].subfield.display_name | Epidemiology |
| topics[0].display_name | Congenital Heart Disease Studies |
| topics[1].id | https://openalex.org/T11677 |
| topics[1].field.id | https://openalex.org/fields/13 |
| topics[1].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[1].score | 0.996399998664856 |
| topics[1].domain.id | https://openalex.org/domains/1 |
| topics[1].domain.display_name | Life Sciences |
| topics[1].subfield.id | https://openalex.org/subfields/1312 |
| topics[1].subfield.display_name | Molecular Biology |
| topics[1].display_name | Congenital heart defects research |
| topics[2].id | https://openalex.org/T10301 |
| topics[2].field.id | https://openalex.org/fields/13 |
| topics[2].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[2].score | 0.9703999757766724 |
| topics[2].domain.id | https://openalex.org/domains/1 |
| topics[2].domain.display_name | Life Sciences |
| topics[2].subfield.id | https://openalex.org/subfields/1312 |
| topics[2].subfield.display_name | Molecular Biology |
| topics[2].display_name | Mitochondrial Function and Pathology |
| is_xpac | False |
| apc_list | |
| apc_paid | |
| concepts[0].id | https://openalex.org/C39133596 |
| concepts[0].level | 2 |
| concepts[0].score | 0.7648138999938965 |
| concepts[0].wikidata | https://www.wikidata.org/wiki/Q5165683 |
| concepts[0].display_name | Contractility |
| concepts[1].id | https://openalex.org/C2779545397 |
| concepts[1].level | 3 |
| concepts[1].score | 0.48922228813171387 |
| concepts[1].wikidata | https://www.wikidata.org/wiki/Q1641399 |
| concepts[1].display_name | Hypoplastic left heart syndrome |
| concepts[2].id | https://openalex.org/C104317684 |
| concepts[2].level | 2 |
| concepts[2].score | 0.45383667945861816 |
| concepts[2].wikidata | https://www.wikidata.org/wiki/Q7187 |
| concepts[2].display_name | Gene |
| concepts[3].id | https://openalex.org/C126322002 |
| concepts[3].level | 1 |
| concepts[3].score | 0.42797911167144775 |
| concepts[3].wikidata | https://www.wikidata.org/wiki/Q11180 |
| concepts[3].display_name | Internal medicine |
| concepts[4].id | https://openalex.org/C164705383 |
| concepts[4].level | 1 |
| concepts[4].score | 0.39092376828193665 |
| concepts[4].wikidata | https://www.wikidata.org/wiki/Q10379 |
| concepts[4].display_name | Cardiology |
| concepts[5].id | https://openalex.org/C86803240 |
| concepts[5].level | 0 |
| concepts[5].score | 0.3858755826950073 |
| concepts[5].wikidata | https://www.wikidata.org/wiki/Q420 |
| concepts[5].display_name | Biology |
| concepts[6].id | https://openalex.org/C95444343 |
| concepts[6].level | 1 |
| concepts[6].score | 0.35397869348526 |
| concepts[6].wikidata | https://www.wikidata.org/wiki/Q7141 |
| concepts[6].display_name | Cell biology |
| concepts[7].id | https://openalex.org/C71924100 |
| concepts[7].level | 0 |
| concepts[7].score | 0.30923065543174744 |
| concepts[7].wikidata | https://www.wikidata.org/wiki/Q11190 |
| concepts[7].display_name | Medicine |
| concepts[8].id | https://openalex.org/C54355233 |
| concepts[8].level | 1 |
| concepts[8].score | 0.26896244287490845 |
| concepts[8].wikidata | https://www.wikidata.org/wiki/Q7162 |
| concepts[8].display_name | Genetics |
| concepts[9].id | https://openalex.org/C2780074459 |
| concepts[9].level | 2 |
| concepts[9].score | 0.06693214178085327 |
| concepts[9].wikidata | https://www.wikidata.org/wiki/Q389735 |
| concepts[9].display_name | Heart disease |
| keywords[0].id | https://openalex.org/keywords/contractility |
| keywords[0].score | 0.7648138999938965 |
| keywords[0].display_name | Contractility |
| keywords[1].id | https://openalex.org/keywords/hypoplastic-left-heart-syndrome |
| keywords[1].score | 0.48922228813171387 |
| keywords[1].display_name | Hypoplastic left heart syndrome |
| keywords[2].id | https://openalex.org/keywords/gene |
| keywords[2].score | 0.45383667945861816 |
| keywords[2].display_name | Gene |
| keywords[3].id | https://openalex.org/keywords/internal-medicine |
| keywords[3].score | 0.42797911167144775 |
| keywords[3].display_name | Internal medicine |
| keywords[4].id | https://openalex.org/keywords/cardiology |
| keywords[4].score | 0.39092376828193665 |
| keywords[4].display_name | Cardiology |
| keywords[5].id | https://openalex.org/keywords/biology |
| keywords[5].score | 0.3858755826950073 |
| keywords[5].display_name | Biology |
| keywords[6].id | https://openalex.org/keywords/cell-biology |
| keywords[6].score | 0.35397869348526 |
| keywords[6].display_name | Cell biology |
| keywords[7].id | https://openalex.org/keywords/medicine |
| keywords[7].score | 0.30923065543174744 |
| keywords[7].display_name | Medicine |
| keywords[8].id | https://openalex.org/keywords/genetics |
| keywords[8].score | 0.26896244287490845 |
| keywords[8].display_name | Genetics |
| keywords[9].id | https://openalex.org/keywords/heart-disease |
| keywords[9].score | 0.06693214178085327 |
| keywords[9].display_name | Heart disease |
| language | en |
| locations[0].id | doi:10.7554/elife.83385.sa2 |
| locations[0].is_oa | True |
| locations[0].source | |
| locations[0].license | cc-by |
| locations[0].pdf_url | |
| locations[0].version | publishedVersion |
| locations[0].raw_type | peer-review |
| locations[0].license_id | https://openalex.org/licenses/cc-by |
| locations[0].is_accepted | True |
| locations[0].is_published | True |
| locations[0].raw_source_name | |
| locations[0].landing_page_url | https://doi.org/10.7554/elife.83385.sa2 |
| indexed_in | crossref |
| authorships[0].author.id | https://openalex.org/A5063024854 |
| authorships[0].author.orcid | https://orcid.org/0000-0003-4789-0971 |
| authorships[0].author.display_name | Katja Birker |
| authorships[0].countries | US |
| authorships[0].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[0].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[0].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[0].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[0].institutions[0].type | nonprofit |
| authorships[0].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[0].institutions[0].country_code | US |
| authorships[0].institutions[0].display_name | Discovery Institute |
| authorships[0].institutions[1].id | https://openalex.org/I188891513 |
| authorships[0].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[0].institutions[1].type | nonprofit |
| authorships[0].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[0].institutions[1].country_code | US |
| authorships[0].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[0].author_position | first |
| authorships[0].raw_author_name | Katja Birker |
| authorships[0].is_corresponding | False |
| authorships[0].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[1].author.id | https://openalex.org/A5013649218 |
| authorships[1].author.orcid | https://orcid.org/0000-0002-6590-9050 |
| authorships[1].author.display_name | Shuchao Ge |
| authorships[1].countries | US |
| authorships[1].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[1].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[1].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[1].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[1].institutions[0].type | nonprofit |
| authorships[1].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[1].institutions[0].country_code | US |
| authorships[1].institutions[0].display_name | Discovery Institute |
| authorships[1].institutions[1].id | https://openalex.org/I188891513 |
| authorships[1].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[1].institutions[1].type | nonprofit |
| authorships[1].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[1].institutions[1].country_code | US |
| authorships[1].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[1].author_position | middle |
| authorships[1].raw_author_name | Shuchao Ge |
| authorships[1].is_corresponding | False |
| authorships[1].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[2].author.id | https://openalex.org/A5054166423 |
| authorships[2].author.orcid | https://orcid.org/0000-0002-7479-1883 |
| authorships[2].author.display_name | Natalie J. Kirkland |
| authorships[2].countries | US |
| authorships[2].affiliations[0].institution_ids | https://openalex.org/I4210089535 |
| authorships[2].affiliations[0].raw_affiliation_string | Department of Bioengineering, Sanford Consortium for Regenerative Medicine, UCSD, School of Medicine |
| authorships[2].institutions[0].id | https://openalex.org/I4210089535 |
| authorships[2].institutions[0].ror | https://ror.org/00cemh325 |
| authorships[2].institutions[0].type | facility |
| authorships[2].institutions[0].lineage | https://openalex.org/I4210089535 |
| authorships[2].institutions[0].country_code | US |
| authorships[2].institutions[0].display_name | Sanford Consortium for Regenerative Medicine |
| authorships[2].author_position | middle |
| authorships[2].raw_author_name | Natalie J Kirkland |
| authorships[2].is_corresponding | False |
| authorships[2].raw_affiliation_strings | Department of Bioengineering, Sanford Consortium for Regenerative Medicine, UCSD, School of Medicine |
| authorships[3].author.id | https://openalex.org/A5018385107 |
| authorships[3].author.orcid | https://orcid.org/0000-0002-4494-8683 |
| authorships[3].author.display_name | Jeanne L. Theis |
| authorships[3].countries | US |
| authorships[3].affiliations[0].institution_ids | https://openalex.org/I2802423016 |
| authorships[3].affiliations[0].raw_affiliation_string | Cardiovascular Genetics Research Laboratory, Mayo Clinic |
| authorships[3].institutions[0].id | https://openalex.org/I2802423016 |
| authorships[3].institutions[0].ror | https://ror.org/02s47w807 |
| authorships[3].institutions[0].type | nonprofit |
| authorships[3].institutions[0].lineage | https://openalex.org/I2802423016 |
| authorships[3].institutions[0].country_code | US |
| authorships[3].institutions[0].display_name | WinnMed |
| authorships[3].author_position | middle |
| authorships[3].raw_author_name | Jeanne L Theis |
| authorships[3].is_corresponding | False |
| authorships[3].raw_affiliation_strings | Cardiovascular Genetics Research Laboratory, Mayo Clinic |
| authorships[4].author.id | https://openalex.org/A5067145283 |
| authorships[4].author.orcid | https://orcid.org/0000-0002-9551-835X |
| authorships[4].author.display_name | James Marchant |
| authorships[4].countries | US |
| authorships[4].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[4].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[4].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[4].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[4].institutions[0].type | nonprofit |
| authorships[4].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[4].institutions[0].country_code | US |
| authorships[4].institutions[0].display_name | Discovery Institute |
| authorships[4].institutions[1].id | https://openalex.org/I188891513 |
| authorships[4].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[4].institutions[1].type | nonprofit |
| authorships[4].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[4].institutions[1].country_code | US |
| authorships[4].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[4].author_position | middle |
| authorships[4].raw_author_name | James Marchant |
| authorships[4].is_corresponding | False |
| authorships[4].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[5].author.id | https://openalex.org/A5005550130 |
| authorships[5].author.orcid | https://orcid.org/0000-0001-5588-3216 |
| authorships[5].author.display_name | Zachary C. Fogarty |
| authorships[5].affiliations[0].raw_affiliation_string | Division of Computational Biology, Department of Quantitative Health Sciences, Mayo Clinic |
| authorships[5].author_position | middle |
| authorships[5].raw_author_name | Zachary C Fogarty |
| authorships[5].is_corresponding | False |
| authorships[5].raw_affiliation_strings | Division of Computational Biology, Department of Quantitative Health Sciences, Mayo Clinic |
| authorships[6].author.id | https://openalex.org/A5023725700 |
| authorships[6].author.orcid | https://orcid.org/0000-0001-9055-758X |
| authorships[6].author.display_name | Maria A. Missinato |
| authorships[6].countries | US |
| authorships[6].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[6].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[6].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[6].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[6].institutions[0].type | nonprofit |
| authorships[6].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[6].institutions[0].country_code | US |
| authorships[6].institutions[0].display_name | Discovery Institute |
| authorships[6].institutions[1].id | https://openalex.org/I188891513 |
| authorships[6].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[6].institutions[1].type | nonprofit |
| authorships[6].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[6].institutions[1].country_code | US |
| authorships[6].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[6].author_position | middle |
| authorships[6].raw_author_name | Maria A Missinato |
| authorships[6].is_corresponding | False |
| authorships[6].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[7].author.id | https://openalex.org/A5035594127 |
| authorships[7].author.orcid | https://orcid.org/0000-0003-3495-3697 |
| authorships[7].author.display_name | Sreehari Kalvakuri |
| authorships[7].countries | US |
| authorships[7].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[7].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[7].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[7].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[7].institutions[0].type | nonprofit |
| authorships[7].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[7].institutions[0].country_code | US |
| authorships[7].institutions[0].display_name | Discovery Institute |
| authorships[7].institutions[1].id | https://openalex.org/I188891513 |
| authorships[7].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[7].institutions[1].type | nonprofit |
| authorships[7].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[7].institutions[1].country_code | US |
| authorships[7].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[7].author_position | middle |
| authorships[7].raw_author_name | Sreehari Kalvakuri |
| authorships[7].is_corresponding | False |
| authorships[7].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[8].author.id | https://openalex.org/A5021669433 |
| authorships[8].author.orcid | https://orcid.org/0000-0001-7215-3821 |
| authorships[8].author.display_name | Paul Grossfeld |
| authorships[8].affiliations[0].raw_affiliation_string | Department of Pediatrics, UCSD School of Medicine, La Jolla, Rady’s Hospital MC 5004 |
| authorships[8].author_position | middle |
| authorships[8].raw_author_name | Paul Grossfeld |
| authorships[8].is_corresponding | False |
| authorships[8].raw_affiliation_strings | Department of Pediatrics, UCSD School of Medicine, La Jolla, Rady’s Hospital MC 5004 |
| authorships[9].author.id | https://openalex.org/A5039405858 |
| authorships[9].author.orcid | https://orcid.org/0000-0003-1642-5380 |
| authorships[9].author.display_name | Adam J. Engler |
| authorships[9].countries | US |
| authorships[9].affiliations[0].institution_ids | https://openalex.org/I4210089535 |
| authorships[9].affiliations[0].raw_affiliation_string | Department of Bioengineering, Sanford Consortium for Regenerative Medicine, UCSD, School of Medicine |
| authorships[9].institutions[0].id | https://openalex.org/I4210089535 |
| authorships[9].institutions[0].ror | https://ror.org/00cemh325 |
| authorships[9].institutions[0].type | facility |
| authorships[9].institutions[0].lineage | https://openalex.org/I4210089535 |
| authorships[9].institutions[0].country_code | US |
| authorships[9].institutions[0].display_name | Sanford Consortium for Regenerative Medicine |
| authorships[9].author_position | middle |
| authorships[9].raw_author_name | Adam J Engler |
| authorships[9].is_corresponding | False |
| authorships[9].raw_affiliation_strings | Department of Bioengineering, Sanford Consortium for Regenerative Medicine, UCSD, School of Medicine |
| authorships[10].author.id | https://openalex.org/A5087552055 |
| authorships[10].author.orcid | https://orcid.org/0000-0003-2593-0119 |
| authorships[10].author.display_name | Karen Ocorr |
| authorships[10].countries | US |
| authorships[10].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[10].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[10].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[10].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[10].institutions[0].type | nonprofit |
| authorships[10].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[10].institutions[0].country_code | US |
| authorships[10].institutions[0].display_name | Discovery Institute |
| authorships[10].institutions[1].id | https://openalex.org/I188891513 |
| authorships[10].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[10].institutions[1].type | nonprofit |
| authorships[10].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[10].institutions[1].country_code | US |
| authorships[10].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[10].author_position | middle |
| authorships[10].raw_author_name | Karen Ocorr |
| authorships[10].is_corresponding | False |
| authorships[10].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[11].author.id | https://openalex.org/A5085716721 |
| authorships[11].author.orcid | https://orcid.org/0000-0002-3862-7023 |
| authorships[11].author.display_name | Timothy J. Nelson |
| authorships[11].countries | US |
| authorships[11].affiliations[0].institution_ids | https://openalex.org/I4210125099 |
| authorships[11].affiliations[0].raw_affiliation_string | Center for Regenerative Medicine, Division of Pediatric Cardiology, Department of Pediatric and Adolescent Medicine, Division of General Internal Medicine, Department of Molecular and Pharmacology and Experimental Therapeutics, Mayo Clinic |
| authorships[11].institutions[0].id | https://openalex.org/I4210125099 |
| authorships[11].institutions[0].ror | https://ror.org/03jp40720 |
| authorships[11].institutions[0].type | healthcare |
| authorships[11].institutions[0].lineage | https://openalex.org/I1330342723, https://openalex.org/I4210125099 |
| authorships[11].institutions[0].country_code | US |
| authorships[11].institutions[0].display_name | Mayo Clinic in Arizona |
| authorships[11].author_position | middle |
| authorships[11].raw_author_name | Timothy J Nelson |
| authorships[11].is_corresponding | False |
| authorships[11].raw_affiliation_strings | Center for Regenerative Medicine, Division of Pediatric Cardiology, Department of Pediatric and Adolescent Medicine, Division of General Internal Medicine, Department of Molecular and Pharmacology and Experimental Therapeutics, Mayo Clinic |
| authorships[12].author.id | https://openalex.org/A5079175915 |
| authorships[12].author.orcid | https://orcid.org/0000-0001-8489-0570 |
| authorships[12].author.display_name | Alexandre R. Colas |
| authorships[12].countries | US |
| authorships[12].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[12].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[12].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[12].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[12].institutions[0].type | nonprofit |
| authorships[12].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[12].institutions[0].country_code | US |
| authorships[12].institutions[0].display_name | Discovery Institute |
| authorships[12].institutions[1].id | https://openalex.org/I188891513 |
| authorships[12].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[12].institutions[1].type | nonprofit |
| authorships[12].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[12].institutions[1].country_code | US |
| authorships[12].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[12].author_position | middle |
| authorships[12].raw_author_name | Alexandre R Colas |
| authorships[12].is_corresponding | False |
| authorships[12].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[13].author.id | https://openalex.org/A5064878571 |
| authorships[13].author.orcid | https://orcid.org/0000-0003-2716-9423 |
| authorships[13].author.display_name | Timothy M. Olson |
| authorships[13].affiliations[0].raw_affiliation_string | Department of Cardiovascular Medicine, Division of Pediatric Cardiology, Department of Pediatric & Adolescent Medicine, Cardiovascular Genetics Research Laboratory, Mayo Clinic |
| authorships[13].author_position | middle |
| authorships[13].raw_author_name | Timothy M Olson |
| authorships[13].is_corresponding | False |
| authorships[13].raw_affiliation_strings | Department of Cardiovascular Medicine, Division of Pediatric Cardiology, Department of Pediatric & Adolescent Medicine, Cardiovascular Genetics Research Laboratory, Mayo Clinic |
| authorships[14].author.id | https://openalex.org/A5087686332 |
| authorships[14].author.orcid | https://orcid.org/0000-0002-8303-3531 |
| authorships[14].author.display_name | Georg Vogler |
| authorships[14].countries | US |
| authorships[14].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[14].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[14].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[14].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[14].institutions[0].type | nonprofit |
| authorships[14].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[14].institutions[0].country_code | US |
| authorships[14].institutions[0].display_name | Discovery Institute |
| authorships[14].institutions[1].id | https://openalex.org/I188891513 |
| authorships[14].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[14].institutions[1].type | nonprofit |
| authorships[14].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[14].institutions[1].country_code | US |
| authorships[14].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[14].author_position | middle |
| authorships[14].raw_author_name | Georg Vogler |
| authorships[14].is_corresponding | False |
| authorships[14].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[15].author.id | https://openalex.org/A5106441010 |
| authorships[15].author.orcid | https://orcid.org/0000-0001-9087-1210 |
| authorships[15].author.display_name | Rolf Bodmer |
| authorships[15].countries | US |
| authorships[15].affiliations[0].institution_ids | https://openalex.org/I188891513, https://openalex.org/I2799424032 |
| authorships[15].affiliations[0].raw_affiliation_string | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| authorships[15].institutions[0].id | https://openalex.org/I2799424032 |
| authorships[15].institutions[0].ror | https://ror.org/05t8s3y29 |
| authorships[15].institutions[0].type | nonprofit |
| authorships[15].institutions[0].lineage | https://openalex.org/I2799424032 |
| authorships[15].institutions[0].country_code | US |
| authorships[15].institutions[0].display_name | Discovery Institute |
| authorships[15].institutions[1].id | https://openalex.org/I188891513 |
| authorships[15].institutions[1].ror | https://ror.org/03m1g2s55 |
| authorships[15].institutions[1].type | nonprofit |
| authorships[15].institutions[1].lineage | https://openalex.org/I188891513 |
| authorships[15].institutions[1].country_code | US |
| authorships[15].institutions[1].display_name | Sanford Burnham Prebys Medical Discovery Institute |
| authorships[15].author_position | last |
| authorships[15].raw_author_name | Rolf Bodmer |
| authorships[15].is_corresponding | False |
| authorships[15].raw_affiliation_strings | Development, Aging and Regeneration Program, Center for Genetic Disorders & Aging Research, Sanford Burnham Prebys Medical Discovery Institute |
| has_content.pdf | False |
| has_content.grobid_xml | False |
| is_paratext | False |
| open_access.is_oa | True |
| open_access.oa_url | https://doi.org/10.7554/elife.83385.sa2 |
| open_access.oa_status | gold |
| open_access.any_repository_has_fulltext | False |
| created_date | 2025-10-10T00:00:00 |
| display_name | Author response: Mitochondrial MICOS complex genes, implicated in hypoplastic left heart syndrome, maintain cardiac contractility and actomyosin integrity |
| has_fulltext | False |
| is_retracted | False |
| updated_date | 2025-11-06T03:46:38.306776 |
| primary_topic.id | https://openalex.org/T10300 |
| primary_topic.field.id | https://openalex.org/fields/27 |
| primary_topic.field.display_name | Medicine |
| primary_topic.score | 0.9987999796867371 |
| primary_topic.domain.id | https://openalex.org/domains/4 |
| primary_topic.domain.display_name | Health Sciences |
| primary_topic.subfield.id | https://openalex.org/subfields/2713 |
| primary_topic.subfield.display_name | Epidemiology |
| primary_topic.display_name | Congenital Heart Disease Studies |
| related_works | https://openalex.org/W4387497383, https://openalex.org/W3183948672, https://openalex.org/W3173606202, https://openalex.org/W3110381201, https://openalex.org/W2948807893, https://openalex.org/W2935909890, https://openalex.org/W2778153218, https://openalex.org/W2758277628, https://openalex.org/W1531601525, https://openalex.org/W4210769129 |
| cited_by_count | 0 |
| locations_count | 1 |
| best_oa_location.id | doi:10.7554/elife.83385.sa2 |
| best_oa_location.is_oa | True |
| best_oa_location.source | |
| best_oa_location.license | cc-by |
| best_oa_location.pdf_url | |
| best_oa_location.version | publishedVersion |
| best_oa_location.raw_type | peer-review |
| best_oa_location.license_id | https://openalex.org/licenses/cc-by |
| best_oa_location.is_accepted | True |
| best_oa_location.is_published | True |
| best_oa_location.raw_source_name | |
| best_oa_location.landing_page_url | https://doi.org/10.7554/elife.83385.sa2 |
| primary_location.id | doi:10.7554/elife.83385.sa2 |
| primary_location.is_oa | True |
| primary_location.source | |
| primary_location.license | cc-by |
| primary_location.pdf_url | |
| primary_location.version | publishedVersion |
| primary_location.raw_type | peer-review |
| primary_location.license_id | https://openalex.org/licenses/cc-by |
| primary_location.is_accepted | True |
| primary_location.is_published | True |
| primary_location.raw_source_name | |
| primary_location.landing_page_url | https://doi.org/10.7554/elife.83385.sa2 |
| publication_date | 2023-06-26 |
| publication_year | 2023 |
| referenced_works_count | 0 |
| abstract_inverted_index.1 | 1362, 1364, 1525, 2585, 2618, 2802, 3066, 3655, 3930, 3938 |
| abstract_inverted_index.2 | 2475, 3588 |
| abstract_inverted_index.9 | 104, 1417, 1528 |
| abstract_inverted_index.A | 1264, 2182, 2741 |
| abstract_inverted_index.B | 1139 |
| abstract_inverted_index.D | 2392 |
| abstract_inverted_index.H | 1237, 1303, 1424, 1535, 2297 |
| abstract_inverted_index.X | 1663 |
| abstract_inverted_index.a | 36, 43, 96, 270, 385, 462, 518, 562, 679, 730, 803, 811, 912, 938, 955, 1164, 1245, 1339, 1600, 1673, 1698, 1713, 1775, 1890, 1995, 2036, 2136, 2208, 2306, 2534, 2544, 2595, 2817, 3711, 3717, 3779, 3964, 4006, 4029, 4134, 4146 |
| abstract_inverted_index.(a | 2170, 2175 |
| abstract_inverted_index.11 | 1236, 1302, 1388, 1423, 1534, 2296 |
| abstract_inverted_index.19 | 2643, 2656, 2669, 2712, 2737, 2757, 2775 |
| abstract_inverted_index.1A | 1498, 4017 |
| abstract_inverted_index.20 | 1678, 2622, 3231, 3673 |
| abstract_inverted_index.29 | 2429, 2576, 2650, 2661, 2674, 2718, 2763, 2828 |
| abstract_inverted_index.2C | 2454 |
| abstract_inverted_index.2E | 2722 |
| abstract_inverted_index.50 | 2566 |
| abstract_inverted_index.6) | 989 |
| abstract_inverted_index.60 | 199 |
| abstract_inverted_index.63 | 1662 |
| abstract_inverted_index.E3 | 231, 1189 |
| abstract_inverted_index.H. | 1389 |
| abstract_inverted_index.Hu | 1848 |
| abstract_inverted_index.II | 1184 |
| abstract_inverted_index.If | 488, 3270 |
| abstract_inverted_index.In | 280, 1198, 1318, 2985, 3925, 4038, 4060, 4107, 4229 |
| abstract_inverted_index.It | 1036 |
| abstract_inverted_index.KD | 154, 217, 1118, 1159, 1876, 1933, 1983, 2031, 2060, 2067, 2151, 2230, 2248, 2405, 2436, 2458, 2709, 2715, 2728, 2744, 2773, 2877, 2943, 3061, 3079, 3113, 3164, 3272, 3329, 3349, 3357, 3393, 3460, 3495, 3528, 3548, 3697, 3730, 3764, 3773, 3802, 3893, 3954, 3995, 4011, 4023, 4045, 4054, 4067, 4100, 4112, 4162, 4200, 4210 |
| abstract_inverted_index.Of | 113, 1980 |
| abstract_inverted_index.PC | 2203 |
| abstract_inverted_index.To | 509, 1799, 2051, 2133, 2688, 2777, 2888, 3309, 3908 |
| abstract_inverted_index.We | 62, 1084, 1142, 2261, 2385, 2423, 3468, 3541, 3850, 3952 |
| abstract_inverted_index.Xu | 2397 |
| abstract_inverted_index.Ye | 638 |
| abstract_inverted_index.an | 92, 192, 886, 930, 1012, 1665, 1745, 3579 |
| abstract_inverted_index.as | 630, 791, 855, 899, 1098, 1100, 1107, 1124, 1126, 1131, 1210, 1772, 1969, 2115, 2117, 2141, 2515, 3131, 3344, 3346, 3984 |
| abstract_inverted_index.at | 569, 2428, 2575, 2584, 2617, 2641, 2649, 2663, 2711, 2717, 2736, 2756, 2762, 2790, 2827, 2847, 2907, 2912, 2969, 3654, 3698, 3760, 3836, 3972, 3977, 4068, 4114, 4170 |
| abstract_inverted_index.be | 413, 1742 |
| abstract_inverted_index.by | 6, 152, 313, 327, 415, 579, 1259, 1396, 2517, 3738, 3986, 4160 |
| abstract_inverted_index.et | 285, 422, 426, 430, 434, 473, 477, 485, 506, 584, 588, 592, 635, 639, 643, 647, 651, 655, 661, 777, 840, 882, 908, 1058, 1354, 1501, 1730, 1780, 1849, 1914, 2042, 2046, 2130, 2320, 2394, 2398, 3300, 3304 |
| abstract_inverted_index.go | 323 |
| abstract_inverted_index.he | 1322 |
| abstract_inverted_index.if | 2266, 2968 |
| abstract_inverted_index.in | 81, 125, 171, 187, 212, 220, 250, 289, 307, 331, 358, 402, 722, 806, 851, 935, 980, 1015, 1018, 1021, 1052, 1067, 1170, 1290, 1375, 1443, 1466, 1554, 1577, 1605, 1687, 1744, 1774, 1787, 1793, 1808, 1869, 1901, 1964, 2021, 2038, 2062, 2101, 2118, 2157, 2185, 2257, 2308, 2329, 2371, 2406, 2461, 2513, 2598, 2753, 2820, 2861, 2883, 2919, 2937, 2948, 2955, 2962, 3043, 3059, 3087, 3106, 3205, 3277, 3293, 3364, 3380, 3387, 3400, 3425, 3436, 3442, 3458, 3477, 3496, 3549, 3601, 3611, 3664, 3708, 3713, 3741, 3769, 3778, 3798, 3847, 3916, 3961, 4042, 4077 |
| abstract_inverted_index.is | 35, 60, 265, 347, 364, 384, 461, 517, 561, 576, 613, 678, 706, 1011, 1027, 1037, 1050, 1073, 1077, 1238, 1492, 1889, 1897, 2466, 2482, 2838, 2885, 2953, 3046, 3530, 3576, 3834 |
| abstract_inverted_index.it | 1888, 3420 |
| abstract_inverted_index.no | 2234, 2751, 3746, 3749, 3795 |
| abstract_inverted_index.of | 50, 88, 95, 118, 132, 155, 159, 218, 225, 247, 254, 261, 273, 355, 392, 441, 453, 464, 564, 571, 603, 607, 674, 712, 715, 733, 760, 820, 848, 863, 914, 916, 933, 940, 985, 990, 1119, 1148, 1160, 1239, 1244, 1269, 1284, 1308, 1373, 1385, 1416, 1473, 1499, 1527, 1591, 1631, 1647, 1667, 1669, 1677, 1735, 1764, 1769, 1835, 1837, 1934, 1955, 1997, 2058, 2068, 2104, 2152, 2231, 2282, 2419, 2459, 2511, 2537, 2562, 2587, 2620, 2628, 2694, 2702, 2705, 2729, 2804, 2857, 2864, 2929, 2941, 3015, 3031, 3054, 3093, 3114, 3137, 3147, 3153, 3165, 3252, 3317, 3330, 3358, 3394, 3396, 3431, 3578, 3607, 3641, 3647, 3657, 3687, 3838, 3882, 3912, 3943, 3958, 3996, 4028, 4133, 4149, 4168 |
| abstract_inverted_index.on | 68, 1431, 1542, 1612, 1622, 1759, 2236, 2275, 3704, 4173, 4232 |
| abstract_inverted_index.or | 189, 234, 418, 448, 495, 1461, 1572, 1603, 1727, 2173, 2204, 2445, 2667, 2795, 3167, 3179, 3219, 3361, 3438, 3740, 3998, 4079 |
| abstract_inverted_index.to | 58, 73, 100, 149, 267, 269, 322, 334, 339, 398, 412, 528, 540, 556, 682, 690, 696, 801, 950, 1029, 1223, 1274, 1320, 1350, 1459, 1570, 1635, 1711, 1741, 1862, 1926, 2092, 2143, 2246, 2264, 2300, 2401, 2433, 2451, 2469, 2543, 2602, 2660, 2673, 2770, 2788, 3123, 3143, 3189, 3202, 3248, 3265, 3291, 3374, 3464, 3489, 3516, 3533, 3865, 4052, 4098, 4109, 4145 |
| abstract_inverted_index.we | 197, 1811, 2028, 2149, 2312, 2345, 2699, 2893, 3260, 3312, 3446, 3503, 3842, 3920, 3968, 4239 |
| abstract_inverted_index.(A) | 1383, 2495, 2589, 3091, 3627 |
| abstract_inverted_index.(B) | 1414, 2524, 3103, 3629 |
| abstract_inverted_index.(C) | 1471, 2569, 3127, 3635 |
| abstract_inverted_index.(D) | 1504, 2606, 3150, 3638 |
| abstract_inverted_index.(E) | 1508, 2625, 3173, 3644 |
| abstract_inverted_index.(F) | 1513, 2642, 3175, 3650 |
| abstract_inverted_index.(G) | 2648, 3177, 3264 |
| abstract_inverted_index.(H) | 2655, 3180 |
| abstract_inverted_index.(I) | 2668, 3213 |
| abstract_inverted_index.(β | 1193 |
| abstract_inverted_index.11H | 1592 |
| abstract_inverted_index.183 | 69, 941 |
| abstract_inverted_index.1B; | 1847 |
| abstract_inverted_index.75% | 759 |
| abstract_inverted_index.89% | 1668 |
| abstract_inverted_index.91% | 1646 |
| abstract_inverted_index.ATP | 139, 156, 834, 856, 1034, 1082, 1132, 1137, 3275, 3283, 3559, 3566, 3636, 3844, 3861 |
| abstract_inverted_index.D). | 2456 |
| abstract_inverted_index.DNA | 1614 |
| abstract_inverted_index.ETC | 176 |
| abstract_inverted_index.For | 809 |
| abstract_inverted_index.Han | 1913 |
| abstract_inverted_index.KD, | 1091, 3919, 4130 |
| abstract_inverted_index.KD. | 1821, 3418, 3433, 3604 |
| abstract_inverted_index.KDs | 2707 |
| abstract_inverted_index.Liu | 425 |
| abstract_inverted_index.PC, | 2160 |
| abstract_inverted_index.RNA | 226, 1182 |
| abstract_inverted_index.TFs | 2519 |
| abstract_inverted_index.The | 320, 437, 1008, 1063, 1295, 1301, 1390, 1451, 1562, 1655, 1852, 1879, 2870, 2927 |
| abstract_inverted_index.V), | 1135 |
| abstract_inverted_index.WGS | 89, 1608 |
| abstract_inverted_index.Xie | 1500 |
| abstract_inverted_index.Yin | 2393 |
| abstract_inverted_index.add | 287, 296 |
| abstract_inverted_index.age | 2805 |
| abstract_inverted_index.al. | 286 |
| abstract_inverted_index.all | 1256, 1367, 2970, 3069, 3539, 3591 |
| abstract_inverted_index.and | 3, 15, 25, 54, 135, 141, 175, 258, 275, 291, 295, 317, 351, 366, 445, 450, 457, 481, 525, 534, 544, 621, 623, 700, 718, 737, 745, 749, 753, 769, 773, 780, 784, 836, 872, 892, 903, 926, 970, 996, 1005, 1033, 1040, 1045, 1070, 1095, 1109, 1140, 1201, 1214, 1218, 1220, 1242, 1251, 1261, 1280, 1282, 1286, 1329, 1338, 1358, 1378, 1408, 1463, 1484, 1512, 1574, 1588, 1640, 1664, 1766, 1797, 1818, 1857, 1893, 1905, 1910, 1949, 1976, 2109, 2180, 2217, 2278, 2291, 2323, 2340, 2367, 2455, 2491, 2500, 2522, 2549, 2558, 2578, 2582, 2613, 2658, 2671, 2867, 2915, 2988, 2999, 3001, 3008, 3040, 3083, 3120, 3186, 3289, 3295, 3321, 3335, 3453, 3456, 3473, 3508, 3555, 3565, 3583, 3609, 3643, 3661, 3691, 3695, 3829, 3860, 3879, 3902, 3935, 3949, 3955, 4018, 4048, 4105, 4152, 4192, 4198, 4202, 4235, 4248 |
| abstract_inverted_index.any | 2446, 3395 |
| abstract_inverted_index.are | 568, 625, 726, 3485, 3680 |
| abstract_inverted_index.but | 47, 1076, 2199, 2227, 2375, 2438, 2845, 3025, 3342, 3370, 3536, 3707, 3797, 3975, 4092 |
| abstract_inverted_index.can | 468 |
| abstract_inverted_index.did | 2376, 2439, 2592, 3018, 3371, 3812, 4071, 4093, 4176, 4221 |
| abstract_inverted_index.due | 1273, 2091, 2245, 3463, 3488, 4097, 4144 |
| abstract_inverted_index.fly | 214, 766, 1966 |
| abstract_inverted_index.for | 195, 207, 390, 514, 632, 708, 1080, 1205, 1212, 1681, 1791, 1866, 1884, 1960, 2078, 2139, 2255, 2272, 2350, 2361, 2484, 2551, 2645, 2652, 2723, 2850, 2890, 3049, 3170, 3210, 3626, 4005 |
| abstract_inverted_index.had | 830, 1298, 1305, 1653, 1697, 1805, 1843, 2083, 2220, 2233, 2731, 3701, 3745, 3794, 4001 |
| abstract_inverted_index.has | 409, 667, 2268 |
| abstract_inverted_index.his | 1249 |
| abstract_inverted_index.how | 692, 2053 |
| abstract_inverted_index.is, | 2088 |
| abstract_inverted_index.its | 1379, 2273, 4174 |
| abstract_inverted_index.may | 694, 2032 |
| abstract_inverted_index.new | 297 |
| abstract_inverted_index.not | 489, 503, 626, 1074, 1753, 2200, 2377, 2440, 2467, 2593, 2839, 3019, 3238, 3343, 3372, 3404, 3531, 3813, 3976, 4072, 4177, 4222 |
| abstract_inverted_index.one | 1403 |
| abstract_inverted_index.our | 48 |
| abstract_inverted_index.out | 1599, 1611, 1642, 1834 |
| abstract_inverted_index.per | 406 |
| abstract_inverted_index.see | 1366, 2178, 2720, 3068, 3590 |
| abstract_inverted_index.set | 732 |
| abstract_inverted_index.six | 1707, 1734 |
| abstract_inverted_index.son | 1404 |
| abstract_inverted_index.the | 51, 82, 160, 167, 244, 281, 353, 362, 367, 370, 403, 454, 511, 541, 548, 557, 600, 605, 671, 697, 716, 723, 781, 795, 917, 974, 991, 1022, 1053, 1278, 1291, 1444, 1448, 1476, 1481, 1489, 1555, 1559, 1606, 1617, 1632, 1636, 1648, 1659, 1670, 1688, 1705, 1717, 1736, 1748, 1760, 1802, 1809, 1813, 1823, 1838, 1902, 1928, 1956, 1965, 1985, 2055, 2119, 2192, 2202, 2221, 2241, 2295, 2372, 2387, 2407, 2462, 2563, 2691, 2703, 2771, 2880, 2908, 2913, 2923, 2938, 3021, 3029, 3036, 3050, 3055, 3088, 3249, 3281, 3314, 3376, 3406, 3429, 3449, 3499, 3517, 3526, 3669, 3809, 3910, 3941, 3944, 4021, 4043, 4074, 4095, 4183, 4252 |
| abstract_inverted_index.two | 1252, 1409 |
| abstract_inverted_index.use | 2701 |
| abstract_inverted_index.via | 1877, 2035 |
| abstract_inverted_index.was | 1271, 1323, 1609, 1661, 1860, 1882, 2005, 2018, 2099, 2188, 2327, 2608, 2767, 2809, 2934, 2965, 2979, 2991, 3004, 3268, 3462, 3752, 3804, 4046, 4123, 4157 |
| abstract_inverted_index.who | 567, 824 |
| abstract_inverted_index.yet | 627 |
| abstract_inverted_index.°C | 2430, 2577, 2644, 2651, 2662, 2675, 2738, 2758, 2764, 2829, 3700, 3728, 4070, 4116, 4172 |
| abstract_inverted_index.µm | 2567, 2623, 3232, 3674 |
| abstract_inverted_index.β. | 1141 |
| abstract_inverted_index.(FS) | 1516 |
| abstract_inverted_index.(Han | 1909 |
| abstract_inverted_index.(KD) | 117, 984 |
| abstract_inverted_index.(Liu | 907 |
| abstract_inverted_index.(Ott | 1057 |
| abstract_inverted_index.(all | 2553 |
| abstract_inverted_index.(see | 2107, 2747 |
| abstract_inverted_index.1A). | 1294, 3327 |
| abstract_inverted_index.1B). | 1979 |
| abstract_inverted_index.1C). | 1918, 3391, 3724, 4037 |
| abstract_inverted_index.1M). | 2252 |
| abstract_inverted_index.2A). | 2343, 4059 |
| abstract_inverted_index.2B). | 2384 |
| abstract_inverted_index.2E). | 2812 |
| abstract_inverted_index.2F), | 2760 |
| abstract_inverted_index.2F). | 2740 |
| abstract_inverted_index.5’ | 1721 |
| abstract_inverted_index.<1%, | 1343 |
| abstract_inverted_index.Bier | 748 |
| abstract_inverted_index.Both | 2215 |
| abstract_inverted_index.CBs, | 2514 |
| abstract_inverted_index.Data | 17 |
| abstract_inverted_index.Duox | 2226, 2232 |
| abstract_inverted_index.Fink | 2129 |
| abstract_inverted_index.Five | 178 |
| abstract_inverted_index.Full | 0 |
| abstract_inverted_index.H15, | 2339, 2521 |
| abstract_inverted_index.HLHS | 59, 70, 97, 180, 274, 312, 332, 400, 408, 442, 470, 501, 565, 633, 687, 807, 821, 864, 887, 922, 942, 977, 1224, 1376, 1406, 1437, 1548, 1689, 1784, 1803, 1840, 2284 |
| abstract_inverted_index.II), | 228 |
| abstract_inverted_index.List | 1415, 1526 |
| abstract_inverted_index.Mef2 | 2165, 2362, 2552 |
| abstract_inverted_index.Open | 1370, 2478, 3072, 3594 |
| abstract_inverted_index.RNAi | 1820, 2076, 2743 |
| abstract_inverted_index.SEM. | 2687, 3230 |
| abstract_inverted_index.Side | 5 |
| abstract_inverted_index.Slit | 2368, 2559 |
| abstract_inverted_index.This | 843, 2097, 2834, 3522 |
| abstract_inverted_index.UMAP | 2496 |
| abstract_inverted_index.Ugur | 776 |
| abstract_inverted_index.WGS. | 1263 |
| abstract_inverted_index.Yagi | 429 |
| abstract_inverted_index.age. | 2588, 3658 |
| abstract_inverted_index.age; | 2621 |
| abstract_inverted_index.aims | 526 |
| abstract_inverted_index.al., | 423, 427, 431, 435, 474, 478, 486, 507, 585, 589, 593, 636, 640, 644, 648, 652, 656, 662, 778, 841, 883, 909, 1059, 1355, 1502, 1731, 1781, 1850, 1915, 2043, 2047, 2131, 2321, 2395, 2399, 3301, 3305 |
| abstract_inverted_index.also | 1143, 1297, 1739, 2082, 2162, 2846, 2980, 3874, 4158 |
| abstract_inverted_index.bars | 2685, 3228 |
| abstract_inverted_index.been | 410, 799, 1348 |
| abstract_inverted_index.body | 1340 |
| abstract_inverted_index.born | 99, 499 |
| abstract_inverted_index.both | 2814 |
| abstract_inverted_index.capt | 1935 |
| abstract_inverted_index.cell | 1153 |
| abstract_inverted_index.core | 731 |
| abstract_inverted_index.data | 4, 90, 368, 928, 1524, 2318, 3889 |
| abstract_inverted_index.down | 1441, 1552 |
| abstract_inverted_index.five | 1618 |
| abstract_inverted_index.from | 91, 204, 324, 818, 1421, 1496, 1517, 1532, 1616, 1755, 2146, 2503, 2533, 2636, 2786, 3076, 3682 |
| abstract_inverted_index.gave | 1984 |
| abstract_inverted_index.gene | 719, 1167, 1785, 2026 |
| abstract_inverted_index.gol, | 1188 |
| abstract_inverted_index.have | 598, 798, 1346, 1712, 2305, 2594 |
| abstract_inverted_index.here | 350 |
| abstract_inverted_index.hg38 | 1649 |
| abstract_inverted_index.high | 2435, 3278 |
| abstract_inverted_index.lack | 913 |
| abstract_inverted_index.lead | 268, 3290 |
| abstract_inverted_index.left | 31, 380, 455, 553 |
| abstract_inverted_index.less | 2825, 2954 |
| abstract_inverted_index.life | 1900, 2432, 2831 |
| abstract_inverted_index.line | 2539, 2541, 2547, 2745, 2774, 3967 |
| abstract_inverted_index.loss | 2536, 3014 |
| abstract_inverted_index.made | 2700 |
| abstract_inverted_index.male | 1246, 3619 |
| abstract_inverted_index.many | 714 |
| abstract_inverted_index.mass | 1341 |
| abstract_inverted_index.mode | 1268 |
| abstract_inverted_index.more | 538 |
| abstract_inverted_index.most | 1986 |
| abstract_inverted_index.need | 681 |
| abstract_inverted_index.next | 2262 |
| abstract_inverted_index.nine | 1692, 1824, 1836 |
| abstract_inverted_index.once | 2676 |
| abstract_inverted_index.only | 2189, 2840, 2992 |
| abstract_inverted_index.over | 1645 |
| abstract_inverted_index.play | 802 |
| abstract_inverted_index.plot | 2502 |
| abstract_inverted_index.rare | 1682 |
| abstract_inverted_index.read | 1675 |
| abstract_inverted_index.risk | 570 |
| abstract_inverted_index.role | 170, 805, 932, 1066, 1868, 2037, 2138, 2307, 3911, 4148 |
| abstract_inverted_index.same | 2222 |
| abstract_inverted_index.side | 7 |
| abstract_inverted_index.site | 995 |
| abstract_inverted_index.size | 452, 906, 3833 |
| abstract_inverted_index.stem | 1152 |
| abstract_inverted_index.such | 790, 854, 1130, 1968 |
| abstract_inverted_index.test | 340, 1800, 1863, 1927, 2052, 2402, 3310 |
| abstract_inverted_index.text | 1, 298 |
| abstract_inverted_index.than | 2002, 2826, 4208 |
| abstract_inverted_index.that | 299, 305, 388, 467, 815, 1026, 1175, 1345, 1625, 1684, 2087, 2124, 2195, 2240, 2325, 2836, 2879, 3013, 3262, 3423, 3448, 3525, 3891, 4254 |
| abstract_inverted_index.they | 624, 693, 1254 |
| abstract_inverted_index.this | 515, 611, 4219 |
| abstract_inverted_index.tin, | 2520 |
| abstract_inverted_index.tinD | 2570, 2590, 2603, 2611, 2614 |
| abstract_inverted_index.tube | 1483 |
| abstract_inverted_index.upon | 1088, 1116, 1158, 2875, 3243, 3546 |
| abstract_inverted_index.used | 1861, 1883, 1925, 2386, 3953 |
| abstract_inverted_index.week | 2586, 2619, 2803, 3656 |
| abstract_inverted_index.well | 348, 1099, 1125, 2116 |
| abstract_inverted_index.were | 78, 147, 967, 1114, 1208, 1255, 1439, 1455, 1464, 1550, 1566, 1575, 1685, 1709, 1738, 1752, 1827, 1924, 2071, 2244, 2359, 2531, 2573, 2580, 2784, 3599, 4185, 4205 |
| abstract_inverted_index.when | 2806, 4126 |
| abstract_inverted_index.will | 502 |
| abstract_inverted_index.with | 42, 107, 166, 210, 222, 491, 500, 610, 890, 1042, 1178, 1247, 1325, 1363, 1405, 1425, 1536, 1990, 2023, 2191, 2201, 2334, 2932, 3065, 3101, 3111, 3161, 3256, 3347, 3411, 3416, 3556, 3587, 3622, 3685, 3743, 3771, 3800, 3822, 3827, 3857, 3906, 3963, 4009, 4065, 4128 |
| abstract_inverted_index.°C, | 2657, 2670, 2713, 3762, 3790, 3974, 3979, 3993 |
| abstract_inverted_index.°C. | 2776 |
| abstract_inverted_index.°C; | 2719 |
| abstract_inverted_index.(CHD) | 41 |
| abstract_inverted_index.(CMs) | 1904 |
| abstract_inverted_index.(Garg | 634 |
| abstract_inverted_index.(PCs) | 1908 |
| abstract_inverted_index.(WGS) | 67, 949 |
| abstract_inverted_index.(Yagi | 1779 |
| abstract_inverted_index.(Yang | 881 |
| abstract_inverted_index.(e.g. | 615, 740 |
| abstract_inverted_index.(five | 1017 |
| abstract_inverted_index.(late | 2528 |
| abstract_inverted_index.(less | 2708 |
| abstract_inverted_index.1993) | 1859 |
| abstract_inverted_index.1995; | 743 |
| abstract_inverted_index.1997; | 2396 |
| abstract_inverted_index.1998) | 2400 |
| abstract_inverted_index.19°C | 2787 |
| abstract_inverted_index.2000; | 586 |
| abstract_inverted_index.2002; | 747 |
| abstract_inverted_index.2004; | 751 |
| abstract_inverted_index.2005; | 637, 1912 |
| abstract_inverted_index.2006; | 1916 |
| abstract_inverted_index.2008; | 590 |
| abstract_inverted_index.2010; | 641, 755, 771 |
| abstract_inverted_index.2011; | 775 |
| abstract_inverted_index.2012; | 1060 |
| abstract_inverted_index.2013. | 1503 |
| abstract_inverted_index.2014; | 645 |
| abstract_inverted_index.2015; | 2044, 2048 |
| abstract_inverted_index.2016; | 653 |
| abstract_inverted_index.2017; | 424, 428, 479, 483, 1356 |
| abstract_inverted_index.2018; | 432 |
| abstract_inverted_index.2020; | 657 |
| abstract_inverted_index.2021; | 659 |
| abstract_inverted_index.29°C | 2789 |
| abstract_inverted_index.99.4% | 1630 |
| abstract_inverted_index.Actin | 1094, 3234 |
| abstract_inverted_index.After | 1638 |
| abstract_inverted_index.Among | 973 |
| abstract_inverted_index.Array | 1594 |
| abstract_inverted_index.CHDs. | 277 |
| abstract_inverted_index.CHDs: | 713 |
| abstract_inverted_index.Cdk12 | 223, 1180 |
| abstract_inverted_index.DIOPT | 1432, 1543 |
| abstract_inverted_index.ETS1, | 618 |
| abstract_inverted_index.HLHS, | 196, 604, 1248, 1321, 1765 |
| abstract_inverted_index.HLHS. | 251, 675, 704, 936 |
| abstract_inverted_index.Hand) | 2341 |
| abstract_inverted_index.Hand. | 2523 |
| abstract_inverted_index.Here, | 937 |
| abstract_inverted_index.Image | 1494 |
| abstract_inverted_index.LRP2, | 620 |
| abstract_inverted_index.MICOS | 120, 168, 308, 992, 1009, 1064, 1121, 1161, 1957, 4256 |
| abstract_inverted_index.MTRR, | 1702 |
| abstract_inverted_index.MYH6, | 619 |
| abstract_inverted_index.Mussa | 480 |
| abstract_inverted_index.Next, | 2344, 3445, 3841 |
| abstract_inverted_index.RNAi. | 1878 |
| abstract_inverted_index.Since | 2015 |
| abstract_inverted_index.Stage | 2525 |
| abstract_inverted_index.Theis | 591, 646, 654, 660 |
| abstract_inverted_index.These | 145, 2080, 3010 |
| abstract_inverted_index.Three | 1695 |
| abstract_inverted_index.Thus, | 2457 |
| abstract_inverted_index.While | 1733 |
| abstract_inverted_index.Whole | 1585 |
| abstract_inverted_index.actin | 134, 1941, 2866, 3287, 3319, 3426, 3475 |
| abstract_inverted_index.adult | 2063, 2121, 2420, 2485, 2493, 2797, 2848 |
| abstract_inverted_index.after | 1315 |
| abstract_inverted_index.along | 2333, 3139, 3155, 3198, 3856 |
| abstract_inverted_index.asset | 1369, 1371, 2477, 2479, 3071, 3073, 3593, 3595 |
| abstract_inverted_index.based | 1430, 1541, 1621, 1758, 4231 |
| abstract_inverted_index.basis | 194 |
| abstract_inverted_index.birth | 386 |
| abstract_inverted_index.blood | 537 |
| abstract_inverted_index.cells | 1907 |
| abstract_inverted_index.chain | 163 |
| abstract_inverted_index.claim | 304 |
| abstract_inverted_index.could | 2304, 3285, 3403 |
| abstract_inverted_index.date, | 510 |
| abstract_inverted_index.depth | 1657, 1676 |
| abstract_inverted_index.early | 2054, 2309, 2492, 2664, 2792, 2796 |
| abstract_inverted_index.equal | 397 |
| abstract_inverted_index.error | 2684, 3227 |
| abstract_inverted_index.exact | 918 |
| abstract_inverted_index.flies | 290, 1923, 2070, 2081, 2106, 2427, 2616, 2639, 2783, 2944, 3080 |
| abstract_inverted_index.force | 838 |
| abstract_inverted_index.found | 721, 1144, 1710, 1740, 2324, 4240 |
| abstract_inverted_index.gene, | 2225 |
| abstract_inverted_index.genes | 106, 203, 257, 608, 689, 717, 764, 849, 960, 978, 1129, 1174, 1419, 1438, 1530, 1549, 1696, 1708, 1751, 1826, 1842, 1865, 3323, 3398, 3690, 3960, 4247, 4258 |
| abstract_inverted_index.given | 670 |
| abstract_inverted_index.great | 680 |
| abstract_inverted_index.heart | 32, 39, 84, 128, 241, 318, 381, 394, 493, 573, 725, 1003, 1227, 1288, 1446, 1482, 1557, 1816, 1870, 2122, 2259, 2276, 2292, 2310, 2373, 2421, 2472, 2697, 2807, 2853, 2872, 3439, 3652, 4233 |
| abstract_inverted_index.human | 292, 762, 1149, 1650, 1839 |
| abstract_inverted_index.iPSCs | 819 |
| abstract_inverted_index.index | 93, 1342, 1386 |
| abstract_inverted_index.inner | 875, 1023, 1952, 3945 |
| abstract_inverted_index.known | 761, 1075 |
| abstract_inverted_index.lacks | 2125 |
| abstract_inverted_index.large | 2183, 3491 |
| abstract_inverted_index.later | 825, 2471 |
| abstract_inverted_index.life, | 2647, 2654 |
| abstract_inverted_index.lines | 2077, 2219 |
| abstract_inverted_index.loss, | 1334 |
| abstract_inverted_index.lungs | 542 |
| abstract_inverted_index.mass, | 1337 |
| abstract_inverted_index.mined | 2313 |
| abstract_inverted_index.model | 356, 889, 1778, 1817 |
| abstract_inverted_index.mouse | 888, 1777 |
| abstract_inverted_index.moved | 2659, 2672 |
| abstract_inverted_index.newly | 685 |
| abstract_inverted_index.novel | 255 |
| abstract_inverted_index.often | 577, 2286 |
| abstract_inverted_index.other | 276, 1120, 1127, 1202, 2147, 2335, 2447, 3296 |
| abstract_inverted_index.outer | 1054 |
| abstract_inverted_index.overt | 2379 |
| abstract_inverted_index.pupal | 2665, 2793, 2843 |
| abstract_inverted_index.rare, | 108, 183, 962 |
| abstract_inverted_index.reads | 1624, 1633 |
| abstract_inverted_index.right | 530, 558, 827, 1309 |
| abstract_inverted_index.ruled | 1598 |
| abstract_inverted_index.seven | 1833, 1929 |
| abstract_inverted_index.share | 787 |
| abstract_inverted_index.shift | 2630 |
| abstract_inverted_index.shown | 2020, 3386, 3835 |
| abstract_inverted_index.since | 2280, 3482 |
| abstract_inverted_index.size, | 870 |
| abstract_inverted_index.small | 614 |
| abstract_inverted_index.stage | 2529 |
| abstract_inverted_index.study | 813, 844, 862 |
| abstract_inverted_index.their | 302, 792, 1829, 1867, 3483, 3490, 3705 |
| abstract_inverted_index.them, | 114 |
| abstract_inverted_index.there | 460, 560, 677 |
| abstract_inverted_index.these | 205, 342, 3397, 3888, 3959 |
| abstract_inverted_index.those | 150 |
| abstract_inverted_index.three | 1173 |
| abstract_inverted_index.trios | 72, 944 |
| abstract_inverted_index.until | 2801 |
| abstract_inverted_index.using | 877, 1447, 1558, 1872, 2073, 2154, 3498, 3668 |
| abstract_inverted_index.where | 959, 1488 |
| abstract_inverted_index.which | 77, 522, 575, 1049, 1105, 1479, 1896, 2007, 2161, 3575, 3768, 3776, 3971, 4121 |
| abstract_inverted_index.white | 1240, 1332 |
| abstract_inverted_index.whole | 64, 328, 946, 2646, 2653 |
| abstract_inverted_index.year. | 407 |
| abstract_inverted_index.years | 1314 |
| abstract_inverted_index.(Brand | 1856 |
| abstract_inverted_index.(CAP1, | 1719 |
| abstract_inverted_index.(CBs), | 2332 |
| abstract_inverted_index.(EDD), | 1507 |
| abstract_inverted_index.(ESD), | 1511 |
| abstract_inverted_index.(ETC), | 164 |
| abstract_inverted_index.(HLHS) | 34, 383 |
| abstract_inverted_index.(Mussa | 543 |
| abstract_inverted_index.(Paige | 839 |
| abstract_inverted_index.(SOHA; | 2128 |
| abstract_inverted_index.(SZT2, | 1701 |
| abstract_inverted_index.(Theis | 472, 969 |
| abstract_inverted_index.(VCMs) | 1157 |
| abstract_inverted_index.(actin | 1936 |
| abstract_inverted_index.(after | 2799 |
| abstract_inverted_index.(early | 2363 |
| abstract_inverted_index.2009). | 2132 |
| abstract_inverted_index.2011). | 1732, 1851 |
| abstract_inverted_index.2015). | 2050, 3951 |
| abstract_inverted_index.2015a; | 475, 649 |
| abstract_inverted_index.2016), | 779 |
| abstract_inverted_index.2016). | 1360, 4154 |
| abstract_inverted_index.2017), | 757 |
| abstract_inverted_index.2017). | 546, 884, 910, 1062 |
| abstract_inverted_index.2018). | 1782 |
| abstract_inverted_index.2019). | 436, 487, 508 |
| abstract_inverted_index.2020). | 842 |
| abstract_inverted_index.2021a) | 2508 |
| abstract_inverted_index.2021b) | 2322 |
| abstract_inverted_index.2022). | 663, 972 |
| abstract_inverted_index.2–4% | 391 |
| abstract_inverted_index.Ahmad, | 756 |
| abstract_inverted_index.Author | 22 |
| abstract_inverted_index.Birker | 284 |
| abstract_inverted_index.Bodmer | 752 |
| abstract_inverted_index.CHCHD3 | 188 |
| abstract_inverted_index.CHCHD6 | 1374 |
| abstract_inverted_index.Cripps | 744 |
| abstract_inverted_index.DGKE), | 1720, 1948 |
| abstract_inverted_index.Family | 1233, 1235 |
| abstract_inverted_index.Figure | 1361, 1497, 1522, 1917, 2111, 2474, 2721, 2748, 3063, 3388, 3585, 4014, 4034, 4084, 4088, 4139, 4214 |
| abstract_inverted_index.First, | 1822, 2299 |
| abstract_inverted_index.Hand), | 2164 |
| abstract_inverted_index.Myosin | 1096, 2902, 2977, 3041, 3457 |
| abstract_inverted_index.Olson, | 746, 971, 1911 |
| abstract_inverted_index.Pandey | 772 |
| abstract_inverted_index.Pcdha9 | 893 |
| abstract_inverted_index.RNF149 | 229, 1186 |
| abstract_inverted_index.SAMM50 | 1043 |
| abstract_inverted_index.SPTBN1 | 235, 1192 |
| abstract_inverted_index.Sap130 | 891 |
| abstract_inverted_index.States | 405 |
| abstract_inverted_index.Theis, | 658 |
| abstract_inverted_index.United | 404 |
| abstract_inverted_index.across | 1658 |
| abstract_inverted_index.and/or | 1225, 1874 |
| abstract_inverted_index.animal | 337 |
| abstract_inverted_index.aorta; | 458 |
| abstract_inverted_index.aortic | 444 |
| abstract_inverted_index.around | 2488 |
| abstract_inverted_index.assays | 2103 |
| abstract_inverted_index.assess | 2301 |
| abstract_inverted_index.author | 26 |
| abstract_inverted_index.begins | 523 |
| abstract_inverted_index.better | 271 |
| abstract_inverted_index.births | 401 |
| abstract_inverted_index.causal | 631 |
| abstract_inverted_index.caused | 239, 414, 2750, 2816, 3337, 3710 |
| abstract_inverted_index.cells, | 1487, 2159 |
| abstract_inverted_index.cohort | 939 |
| abstract_inverted_index.defect | 387 |
| abstract_inverted_index.delay, | 1327 |
| abstract_inverted_index.divert | 552 |
| abstract_inverted_index.dorsal | 2408 |
| abstract_inverted_index.double | 1397, 4044 |
| abstract_inverted_index.driven | 2229, 3163 |
| abstract_inverted_index.driver | 1491, 1881, 2194, 2389, 3501 |
| abstract_inverted_index.during | 2413, 2696, 2842 |
| abstract_inverted_index.effect | 2210, 2235, 3703 |
| abstract_inverted_index.either | 2791, 4053 |
| abstract_inverted_index.factor | 2168 |
| abstract_inverted_index.family | 94, 958, 1387, 1391, 1593, 1619 |
| abstract_inverted_index.father | 1281 |
| abstract_inverted_index.female | 1468, 1519, 1579, 1922, 2638, 3108, 3666 |
| abstract_inverted_index.filmed | 2581 |
| abstract_inverted_index.flies. | 1521, 2605 |
| abstract_inverted_index.genes, | 76, 953, 1123, 3900 |
| abstract_inverted_index.genes. | 1162, 1694, 1931 |
| abstract_inverted_index.genome | 65, 329, 947, 1586, 1652, 1660, 1671 |
| abstract_inverted_index.having | 765 |
| abstract_inverted_index.heart, | 1810 |
| abstract_inverted_index.hearts | 786, 2281, 3110, 3160, 3461, 3667, 3873, 4184 |
| abstract_inverted_index.imaged | 2583 |
| abstract_inverted_index.impact | 2470 |
| abstract_inverted_index.inputs | 2127 |
| abstract_inverted_index.intact | 2105 |
| abstract_inverted_index.intron | 1728 |
| abstract_inverted_index.larval | 2489 |
| abstract_inverted_index.latent | 572, 1226, 1306 |
| abstract_inverted_index.letter | 21, 374 |
| abstract_inverted_index.levels | 131, 2869, 3042, 3254, 3284, 3383 |
| abstract_inverted_index.likely | 44, 245, 1078, 1221, 3047, 3537 |
| abstract_inverted_index.lines) | 1399 |
| abstract_inverted_index.listed | 1454, 1565 |
| abstract_inverted_index.lumen) | 2374 |
| abstract_inverted_index.lumen, | 2564 |
| abstract_inverted_index.mapped | 1634 |
| abstract_inverted_index.matter | 1333 |
| abstract_inverted_index.mitral | 446 |
| abstract_inverted_index.model. | 85 |
| abstract_inverted_index.models | 338 |
| abstract_inverted_index.mother | 1279 |
| abstract_inverted_index.muscle | 1336, 2554 |
| abstract_inverted_index.mutant | 2348 |
| abstract_inverted_index.myosin | 2868 |
| abstract_inverted_index.nature | 1763 |
| abstract_inverted_index.normal | 1299, 3412 |
| abstract_inverted_index.number | 606 |
| abstract_inverted_index.origin | 793 |
| abstract_inverted_index.passed | 1626 |
| abstract_inverted_index.proved | 668 |
| abstract_inverted_index.rather | 2001 |
| abstract_inverted_index.reads, | 1644 |
| abstract_inverted_index.reads. | 1679 |
| abstract_inverted_index.reared | 2424, 2574, 2640 |
| abstract_inverted_index.recent | 812, 924 |
| abstract_inverted_index.reduce | 2033 |
| abstract_inverted_index.region | 1478, 1723 |
| abstract_inverted_index.review | 376 |
| abstract_inverted_index.robust | 2852 |
| abstract_inverted_index.scale. | 2568, 2624, 3233, 3649, 3675 |
| abstract_inverted_index.score. | 1433, 1544 |
| abstract_inverted_index.screen | 1169 |
| abstract_inverted_index.severe | 37, 438, 1002, 1101, 1987, 2862, 3345 |
| abstract_inverted_index.showed | 895 |
| abstract_inverted_index.sought | 2263 |
| abstract_inverted_index.stage) | 1921 |
| abstract_inverted_index.stages | 2415, 2490, 2794, 2798, 2849 |
| abstract_inverted_index.strong | 1891, 2732, 2871, 3250 |
| abstract_inverted_index.system | 1855 |
| abstract_inverted_index.tested | 80, 198, 979 |
| abstract_inverted_index.those, | 1981 |
| abstract_inverted_index.traits | 1344 |
| abstract_inverted_index.weaker | 2742 |
| abstract_inverted_index.within | 794, 1716 |
| abstract_inverted_index.(Alston | 1353 |
| abstract_inverted_index.(Bodmer | 768 |
| abstract_inverted_index.(CHDs), | 396 |
| abstract_inverted_index.(Diacyl | 1945 |
| abstract_inverted_index.(Figure | 1293, 1846, 1978, 2011, 2095, 2211, 2249, 2342, 2383, 2453, 2739, 2759, 2811, 2832, 2925, 2945, 2971, 2983, 2995, 3006, 3324, 3350, 3513, 3520, 3573, 3721, 3754, 3783, 3816, 3867, 3884, 3989, 4012, 4032, 4056, 4082, 4101, 4137, 4163, 4180, 4188, 4212, 4225 |
| abstract_inverted_index.(F–I) | 2632 |
| abstract_inverted_index.(Vogler | 2319 |
| abstract_inverted_index.(tinman | 2390 |
| abstract_inverted_index.(virgin | 2678 |
| abstract_inverted_index.10–12 | 2416 |
| abstract_inverted_index.16–17 | 2354, 2526 |
| abstract_inverted_index.1D–F) | 2749 |
| abstract_inverted_index.2)-Gal4 | 2169 |
| abstract_inverted_index.2015b). | 594 |
| abstract_inverted_index.Another | 861 |
| abstract_inverted_index.Article | 24 |
| abstract_inverted_index.Barron, | 482, 545 |
| abstract_inverted_index.Bodmer, | 750 |
| abstract_inverted_index.CHCHD6. | 190 |
| abstract_inverted_index.Cardiac | 2855, 3074, 3328 |
| abstract_inverted_index.ChchdD1 | 2355 |
| abstract_inverted_index.Crucean | 476 |
| abstract_inverted_index.Despite | 911 |
| abstract_inverted_index.F-actin | 2607, 2930, 2960, 2975, 3039, 3104, 3128, 3154, 3253, 3266, 3294, 3382, 3401, 3409, 3455, 3506, 3608, 3660, 3859, 4040, 4080, 4155, 4203, 4236 |
| abstract_inverted_index.Figures | 2 |
| abstract_inverted_index.Frasch, | 754, 770 |
| abstract_inverted_index.Further | 252 |
| abstract_inverted_index.MBTPS1) | 1703 |
| abstract_inverted_index.McBride | 587 |
| abstract_inverted_index.Metrics | 28 |
| abstract_inverted_index.NKX2-5, | 616 |
| abstract_inverted_index.NOTCH1, | 617 |
| abstract_inverted_index.RNF149, | 1725 |
| abstract_inverted_index.Results | 12, 1232 |
| abstract_inverted_index.Sandri, | 1359 |
| abstract_inverted_index.Similar | 1112 |
| abstract_inverted_index.Tocchi, | 2049 |
| abstract_inverted_index.ability | 321 |
| abstract_inverted_index.absence | 1283 |
| abstract_inverted_index.achieve | 529, 2434 |
| abstract_inverted_index.adapted | 1495 |
| abstract_inverted_index.affects | 2417, 3538 |
| abstract_inverted_index.already | 2735, 4113 |
| abstract_inverted_index.animals | 2022 |
| abstract_inverted_index.average | 1656, 1666 |
| abstract_inverted_index.because | 1887 |
| abstract_inverted_index.between | 1277, 2610, 3581, 4244 |
| abstract_inverted_index.binding | 1937 |
| abstract_inverted_index.cardiac | 138, 153, 345, 439, 465, 496, 1068, 1117, 1216, 1401, 1412, 1795, 1971, 1988, 1998, 2056, 2102, 2197, 2242, 2351, 2380, 2411, 2464, 2486, 2518, 3044, 3244, 3315, 3356, 3518, 3534, 3602, 3895, 3917, 3994 |
| abstract_inverted_index.carried | 1610 |
| abstract_inverted_index.central | 303 |
| abstract_inverted_index.claims. | 371 |
| abstract_inverted_index.closely | 1038 |
| abstract_inverted_index.complex | 121, 309, 1000, 1010, 1014, 1122, 4257 |
| abstract_inverted_index.contact | 994 |
| abstract_inverted_index.control | 1628, 3166 |
| abstract_inverted_index.coupled | 858 |
| abstract_inverted_index.cristae | 173, 901, 997, 1031 |
| abstract_inverted_index.crossed | 2542 |
| abstract_inverted_index.decline | 1307 |
| abstract_inverted_index.defects | 146, 395, 897, 1963, 1989, 2382, 2734, 2874, 3424, 3572, 3598, 3878, 3915 |
| abstract_inverted_index.deliver | 535 |
| abstract_inverted_index.density | 902 |
| abstract_inverted_index.derived | 817, 1420, 1531 |
| abstract_inverted_index.develop | 2287 |
| abstract_inverted_index.digenic | 1776 |
| abstract_inverted_index.disease | 40, 359, 516, 612, 1267, 1289 |
| abstract_inverted_index.diverse | 248 |
| abstract_inverted_index.driver, | 1895, 3672 |
| abstract_inverted_index.driver. | 1450, 1561 |
| abstract_inverted_index.driver; | 2177 |
| abstract_inverted_index.eLife's | 375 |
| abstract_inverted_index.eclosed | 2677 |
| abstract_inverted_index.effects | 2145, 2243, 2274, 3430 |
| abstract_inverted_index.elusive | 669 |
| abstract_inverted_index.embryos | 2349, 2358, 2527 |
| abstract_inverted_index.examine | 2778 |
| abstract_inverted_index.exhibit | 2378, 3081 |
| abstract_inverted_index.express | 2163 |
| abstract_inverted_index.factor) | 2366 |
| abstract_inverted_index.factor, | 2556 |
| abstract_inverted_index.factors | 420, 736, 739, 2337 |
| abstract_inverted_index.failure | 1228 |
| abstract_inverted_index.females | 2579, 3684 |
| abstract_inverted_index.flies), | 2679 |
| abstract_inverted_index.further | 300, 2134, 2689, 3523, 3805, 4206 |
| abstract_inverted_index.fusion) | 1961 |
| abstract_inverted_index.genetic | 52, 208, 259, 710, 3719, 3750, 3781, 3922, 4030, 4135, 4242 |
| abstract_inverted_index.genome. | 1637 |
| abstract_inverted_index.genomic | 1596, 1613 |
| abstract_inverted_index.greater | 2714 |
| abstract_inverted_index.green). | 2565 |
| abstract_inverted_index.hearts, | 1967 |
| abstract_inverted_index.hearts. | 215, 1470, 1581, 3062, 3849 |
| abstract_inverted_index.include | 443 |
| abstract_inverted_index.induced | 1150 |
| abstract_inverted_index.infants | 498 |
| abstract_inverted_index.initial | 1885 |
| abstract_inverted_index.kinase, | 1947 |
| abstract_inverted_index.knocked | 1440, 1551 |
| abstract_inverted_index.leading | 57 |
| abstract_inverted_index.levels, | 140, 1097 |
| abstract_inverted_index.located | 1020, 1051 |
| abstract_inverted_index.mammals | 1016 |
| abstract_inverted_index.marking | 1639 |
| abstract_inverted_index.measure | 1996 |
| abstract_inverted_index.mediate | 311, 3239, 3428 |
| abstract_inverted_index.methods | 16 |
| abstract_inverted_index.minimal | 1674 |
| abstract_inverted_index.muscle, | 2198 |
| abstract_inverted_index.myosin, | 136 |
| abstract_inverted_index.number, | 871 |
| abstract_inverted_index.observe | 2441 |
| abstract_inverted_index.opposed | 2142 |
| abstract_inverted_index.output, | 1972 |
| abstract_inverted_index.overall | 527, 3022 |
| abstract_inverted_index.parents | 102, 1292, 1394 |
| abstract_inverted_index.proband | 98, 1304, 1377, 1422, 1533, 1690 |
| abstract_inverted_index.process | 377 |
| abstract_inverted_index.produce | 344, 3375 |
| abstract_inverted_index.protein | 988, 1954, 2370, 2561, 3095, 3929 |
| abstract_inverted_index.quality | 1627 |
| abstract_inverted_index.reduced | 137, 451, 580, 831, 868, 900, 1086, 1970, 1973, 1992, 2016, 2024, 2084, 2089, 2288, 2442, 2768, 2994, 3082, 3257, 3282, 3313, 3339, 3407, 3450, 3465, 3505, 3731, 3766, 3806, 3858, 3880, 4024, 4207 |
| abstract_inverted_index.related | 1349, 3928 |
| abstract_inverted_index.results | 2860, 3441, 3679 |
| abstract_inverted_index.reveals | 1789 |
| abstract_inverted_index.revised | 282 |
| abstract_inverted_index.samples | 1615 |
| abstract_inverted_index.several | 596, 1313, 2074, 2895, 3318 |
| abstract_inverted_index.shifted | 2785 |
| abstract_inverted_index.showing | 2509 |
| abstract_inverted_index.similar | 148, 3515 |
| abstract_inverted_index.smaller | 904 |
| abstract_inverted_index.spatial | 1875 |
| abstract_inverted_index.stages, | 2064, 2666 |
| abstract_inverted_index.stages. | 2494 |
| abstract_inverted_index.stained | 2360, 2550, 3169, 3209, 3625 |
| abstract_inverted_index.studies | 597 |
| abstract_inverted_index.subunit | 122 |
| abstract_inverted_index.suggest | 929, 2878, 3012 |
| abstract_inverted_index.support | 369 |
| abstract_inverted_index.survive | 504 |
| abstract_inverted_index.system) | 999 |
| abstract_inverted_index.t-test, | 2682, 3224 |
| abstract_inverted_index.treated | 490 |
| abstract_inverted_index.underly | 469 |
| abstract_inverted_index.variant | 1700, 1715 |
| abstract_inverted_index.whereas | 1704, 2761, 3735 |
| abstract_inverted_index.whether | 341, 1801, 2302, 2403 |
| abstract_inverted_index.without | 1400, 1411 |
| abstract_inverted_index.(Altmann | 583 |
| abstract_inverted_index.(Bodmer, | 742 |
| abstract_inverted_index.(Crucean | 421 |
| abstract_inverted_index.(Myocyte | 2166 |
| abstract_inverted_index.(RHBDL2, | 1724 |
| abstract_inverted_index.(Vogler, | 2507 |
| abstract_inverted_index.(complex | 1134, 3553, 3561 |
| abstract_inverted_index.(denoted | 1395 |
| abstract_inverted_index.(sorting | 1044 |
| abstract_inverted_index.(tinman, | 2338 |
| abstract_inverted_index.(uniform | 2497 |
| abstract_inverted_index.1A–C). | 2014 |
| abstract_inverted_index.1D–F), | 2114 |
| abstract_inverted_index.1D–F). | 2096 |
| abstract_inverted_index.1J–L). | 2214 |
| abstract_inverted_index.2G–I). | 2833 |
| abstract_inverted_index.Abstract | 8, 29 |
| abstract_inverted_index.Although | 547, 595 |
| abstract_inverted_index.CELSR1), | 622 |
| abstract_inverted_index.CG7639), | 1048 |
| abstract_inverted_index.CHCHD3/6 | 211, 219 |
| abstract_inverted_index.Chchd3/6 | 986, 1090, 1200, 1792, 1950, 1982, 2030, 2059, 2069, 2153, 2247, 2256, 2267, 2303, 2326, 2347, 2404, 2460, 2480, 2512, 2535, 2545, 2695, 2837, 2859, 2876, 3016, 3060, 3078, 3168, 3245, 3258, 3271, 3348, 3377, 3417, 3432, 3434, 3459, 3494, 3527, 3547, 3603, 3694, 3729, 3772, 3801, 3892, 3907, 3918, 3965, 3999, 4010, 4066, 4099, 4129, 4199, 4209 |
| abstract_inverted_index.ControlA | 1460, 1571 |
| abstract_inverted_index.ControlB | 1462, 1573 |
| abstract_inverted_index.Decision | 20, 373 |
| abstract_inverted_index.Defining | 664 |
| abstract_inverted_index.Dot-Gal4 | 2155 |
| abstract_inverted_index.Download | 1368, 1583, 2476, 3070, 3592 |
| abstract_inverted_index.Editor's | 9, 278 |
| abstract_inverted_index.Finally, | 1163 |
| abstract_inverted_index.Gal4-UAS | 1854 |
| abstract_inverted_index.Moderate | 216 |
| abstract_inverted_index.Nichols, | 774 |
| abstract_inverted_index.Overall, | 361, 676, 3035, 3392 |
| abstract_inverted_index.Pedigree | 1384 |
| abstract_inverted_index.Systolic | 2000 |
| abstract_inverted_index.Temporal | 2253 |
| abstract_inverted_index.Unpaired | 2680, 3222 |
| abstract_inverted_index.accounts | 389 |
| abstract_inverted_index.addition | 1319 |
| abstract_inverted_index.although | 2823, 3808, 4182, 4218 |
| abstract_inverted_index.analyses | 1757 |
| abstract_inverted_index.analysis | 87, 1590 |
| abstract_inverted_index.analyzed | 2346 |
| abstract_inverted_index.ancestry | 1241 |
| abstract_inverted_index.assembly | 1046 |
| abstract_inverted_index.assessed | 2810 |
| abstract_inverted_index.assigned | 1828 |
| abstract_inverted_index.atresia, | 449 |
| abstract_inverted_index.atrophy, | 1331 |
| abstract_inverted_index.cerebral | 1328 |
| abstract_inverted_index.compared | 2601, 4051 |
| abstract_inverted_index.complex, | 1958 |
| abstract_inverted_index.controls | 2452, 3866 |
| abstract_inverted_index.critical | 804, 1211 |
| abstract_inverted_index.damaging | 110, 185, 964 |
| abstract_inverted_index.defects, | 242, 1104, 1402, 2449, 3855 |
| abstract_inverted_index.defects. | 144, 1007, 1413 |
| abstract_inverted_index.deletion | 1602 |
| abstract_inverted_index.diameter | 1506, 1510, 2004 |
| abstract_inverted_index.directly | 539 |
| abstract_inverted_index.driver), | 2172 |
| abstract_inverted_index.drivers, | 2206 |
| abstract_inverted_index.ejection | 581, 1311, 2289 |
| abstract_inverted_index.electron | 161, 859, 879 |
| abstract_inverted_index.emerging | 686 |
| abstract_inverted_index.enhancer | 2167, 2391 |
| abstract_inverted_index.evaluate | 684 |
| abstract_inverted_index.examined | 599, 2894, 3470, 3543 |
| abstract_inverted_index.example, | 810 |
| abstract_inverted_index.excluded | 1754 |
| abstract_inverted_index.expected | 266 |
| abstract_inverted_index.failure, | 574, 829, 2293 |
| abstract_inverted_index.filament | 1942 |
| abstract_inverted_index.flanking | 1485 |
| abstract_inverted_index.fraction | 582, 1312, 2290 |
| abstract_inverted_index.function | 555, 1006, 1798, 1871, 2040, 2260, 2487, 2538, 2808, 3017 |
| abstract_inverted_index.genetic, | 416 |
| abstract_inverted_index.glycerol | 1946 |
| abstract_inverted_index.harbored | 182 |
| abstract_inverted_index.however, | 459, 4020, 4194 |
| abstract_inverted_index.iPSC-CMs | 866 |
| abstract_inverted_index.identify | 74, 951 |
| abstract_inverted_index.includes | 1392, 1480, 2196 |
| abstract_inverted_index.interact | 1177 |
| abstract_inverted_index.involved | 850, 3399 |
| abstract_inverted_index.ligase), | 233, 1191 |
| abstract_inverted_index.limited. | 61 |
| abstract_inverted_index.magenta) | 2557 |
| abstract_inverted_index.maintain | 1030 |
| abstract_inverted_index.manifold | 2498 |
| abstract_inverted_index.measured | 1465, 1576, 3130, 3985 |
| abstract_inverted_index.mediated | 2727 |
| abstract_inverted_index.members, | 1620 |
| abstract_inverted_index.membrane | 1025, 1056, 1953, 3947 |
| abstract_inverted_index.mesoderm | 2409, 2465 |
| abstract_inverted_index.methods; | 2110 |
| abstract_inverted_index.missense | 1699 |
| abstract_inverted_index.modeling | 709 |
| abstract_inverted_index.neuronal | 2126, 2205 |
| abstract_inverted_index.observed | 1085, 1115, 1773, 2100, 2190, 3255, 3504, 3600, 3753, 3852 |
| abstract_inverted_index.operated | 2283 |
| abstract_inverted_index.ortholog | 1381 |
| abstract_inverted_index.overview | 2627 |
| abstract_inverted_index.parents, | 1250 |
| abstract_inverted_index.pathways | 249, 264 |
| abstract_inverted_index.patients | 206, 333, 566, 822, 2285 |
| abstract_inverted_index.preceded | 578 |
| abstract_inverted_index.proband. | 1607, 2298 |
| abstract_inverted_index.probands | 181 |
| abstract_inverted_index.produced | 1962 |
| abstract_inverted_index.programs | 720 |
| abstract_inverted_index.promoter | 1718 |
| abstract_inverted_index.proposed | 411 |
| abstract_inverted_index.protein) | 238 |
| abstract_inverted_index.protein, | 1938 |
| abstract_inverted_index.reducing | 314 |
| abstract_inverted_index.relative | 1458, 1569, 2450, 3142, 3201, 3864 |
| abstract_inverted_index.reported | 814, 1275 |
| abstract_inverted_index.required | 1959, 2841 |
| abstract_inverted_index.response | 23 |
| abstract_inverted_index.resulted | 124, 3363, 3777 |
| abstract_inverted_index.revealed | 845, 867, 1172, 1691, 3569 |
| abstract_inverted_index.sibling, | 1747 |
| abstract_inverted_index.siblings | 1296, 1410 |
| abstract_inverted_index.somewhat | 2824, 4186 |
| abstract_inverted_index.spectrum | 463 |
| abstract_inverted_index.standard | 512 |
| abstract_inverted_index.stenosis | 447 |
| abstract_inverted_index.stronger | 2772 |
| abstract_inverted_index.strongly | 1991, 3037, 4117 |
| abstract_inverted_index.subgroup | 563 |
| abstract_inverted_index.subunits | 158 |
| abstract_inverted_index.suggests | 2008, 2835 |
| abstract_inverted_index.summary, | 1199, 4230 |
| abstract_inverted_index.supports | 301 |
| abstract_inverted_index.surgical | 520, 549, 1230, 1316 |
| abstract_inverted_index.syndrome | 33, 382 |
| abstract_inverted_index.synthase | 157, 1133, 1138, 3560, 3567, 3845, 3862 |
| abstract_inverted_index.systemic | 532 |
| abstract_inverted_index.systolic | 2009, 2093, 4119 |
| abstract_inverted_index.temporal | 1873, 2270, 2692 |
| abstract_inverted_index.tissues, | 2148 |
| abstract_inverted_index.utilized | 1812 |
| abstract_inverted_index.validate | 2135 |
| abstract_inverted_index.variants | 186, 326, 343, 966, 1683, 1737 |
| abstract_inverted_index.version, | 283 |
| abstract_inverted_index.windows, | 2781 |
| abstract_inverted_index.β-Spec, | 1195 |
| abstract_inverted_index.(Bhandari | 2041 |
| abstract_inverted_index.(ChchdD1) | 2540 |
| abstract_inverted_index.(goliath, | 230, 1187 |
| abstract_inverted_index.(secreted | 2369, 2560 |
| abstract_inverted_index.>ControlA | 2571, 2604, 2612 |
| abstract_inverted_index.Candidate | 1783 |
| abstract_inverted_index.Chchd3/6, | 2140, 4249 |
| abstract_inverted_index.Chchd3/6. | 1382, 2079, 3115 |
| abstract_inverted_index.Chchd3/6: | 1179 |
| abstract_inverted_index.Conserved | 1434, 1545 |
| abstract_inverted_index.Filtering | 1680 |
| abstract_inverted_index.Grossfeld | 433, 484 |
| abstract_inverted_index.Kobayashi | 642 |
| abstract_inverted_index.Materials | 14, 2108, 2179 |
| abstract_inverted_index.Orthology | 1429, 1540 |
| abstract_inverted_index.Perrimon, | 1858 |
| abstract_inverted_index.Romanello | 1357 |
| abstract_inverted_index.Schematic | 1472, 2626, 3092 |
| abstract_inverted_index.Spectrin, | 1194 |
| abstract_inverted_index.abdominal | 1477 |
| abstract_inverted_index.assembly. | 177 |
| abstract_inverted_index.candidate | 75, 105, 202, 256, 688, 952, 976, 1166, 1418, 1436, 1529, 1547, 1693, 1750, 1825, 1841, 1864, 1930 |
| abstract_inverted_index.cellular, | 699 |
| abstract_inverted_index.collected | 2532 |
| abstract_inverted_index.comprised | 1243 |
| abstract_inverted_index.correctly | 551 |
| abstract_inverted_index.coverage. | 1654 |
| abstract_inverted_index.dCHCHD3/6 | 123 |
| abstract_inverted_index.decreased | 1335 |
| abstract_inverted_index.developed | 826 |
| abstract_inverted_index.diagnosed | 1324 |
| abstract_inverted_index.diastolic | 2003 |
| abstract_inverted_index.different | 1457, 1568, 2269, 2779 |
| abstract_inverted_index.duplicate | 1643 |
| abstract_inverted_index.eclosion) | 2800 |
| abstract_inverted_index.efficient | 1081 |
| abstract_inverted_index.elav-Gal4 | 2174 |
| abstract_inverted_index.embryonic | 2314, 2414, 2463 |
| abstract_inverted_index.etiology, | 46 |
| abstract_inverted_index.examined, | 2072 |
| abstract_inverted_index.exhibited | 1001, 3875, 4118 |
| abstract_inverted_index.expressed | 1898, 2328 |
| abstract_inverted_index.filtering | 1641 |
| abstract_inverted_index.following | 1229 |
| abstract_inverted_index.function, | 1219, 2279 |
| abstract_inverted_index.function. | 2422, 2473, 2854, 3467 |
| abstract_inverted_index.harboring | 961 |
| abstract_inverted_index.homologs; | 1832 |
| abstract_inverted_index.important | 1079, 1204, 2483 |
| abstract_inverted_index.improved, | 365, 3775 |
| abstract_inverted_index.including | 729, 954, 2294, 2898, 3368 |
| abstract_inverted_index.inductive | 738 |
| abstract_inverted_index.inflicted | 151 |
| abstract_inverted_index.integrity | 316, 3472, 3545 |
| abstract_inverted_index.interacts | 1041 |
| abstract_inverted_index.knockdown | 116, 983, 1786, 2856, 3246, 3435 |
| abstract_inverted_index.malformed | 873 |
| abstract_inverted_index.mammalian | 783 |
| abstract_inverted_index.manifests | 2061 |
| abstract_inverted_index.membranes | 876 |
| abstract_inverted_index.mesoderm) | 2412 |
| abstract_inverted_index.mesoderm. | 796 |
| abstract_inverted_index.methods). | 2181 |
| abstract_inverted_index.molecular | 601 |
| abstract_inverted_index.mutations | 306, 894 |
| abstract_inverted_index.myopathic | 1287 |
| abstract_inverted_index.necessary | 1028 |
| abstract_inverted_index.organisms | 357 |
| abstract_inverted_index.orthologs | 767, 1845 |
| abstract_inverted_index.performed | 63, 2150 |
| abstract_inverted_index.phenotype | 1234, 2057, 2098, 3519, 3529 |
| abstract_inverted_index.predicted | 109, 184, 963, 2223 |
| abstract_inverted_index.processes | 702 |
| abstract_inverted_index.protein). | 1197 |
| abstract_inverted_index.recessive | 1266 |
| abstract_inverted_index.reduction | 2184, 2597, 2752, 2819, 2863, 3242, 3251, 3379, 3712 |
| abstract_inverted_index.reference | 1651 |
| abstract_inverted_index.remaining | 1706 |
| abstract_inverted_index.represent | 2686, 3229 |
| abstract_inverted_index.research. | 360 |
| abstract_inverted_index.resulting | 975 |
| abstract_inverted_index.sarcomere | 315, 3510 |
| abstract_inverted_index.screening | 1886 |
| abstract_inverted_index.siblings, | 1253 |
| abstract_inverted_index.similarly | 2769 |
| abstract_inverted_index.structure | 1004, 1217, 1796, 2277 |
| abstract_inverted_index.surgeries | 494 |
| abstract_inverted_index.synthesis | 857 |
| abstract_inverted_index.tinD-Gal4 | 2388, 2425 |
| abstract_inverted_index.transport | 162 |
| abstract_inverted_index.treatment | 513 |
| abstract_inverted_index.ubiquitin | 232, 1190 |
| abstract_inverted_index.unchanged | 2609 |
| abstract_inverted_index.underwent | 945, 1262 |
| abstract_inverted_index.variants, | 1771 |
| abstract_inverted_index.variants. | 112 |
| abstract_inverted_index.ventricle | 456 |
| abstract_inverted_index.(Grossfeld | 505 |
| abstract_inverted_index.(activator | 224, 1181 |
| abstract_inverted_index.(expressed | 2156 |
| abstract_inverted_index.(iPSC-CM), | 823 |
| abstract_inverted_index.(including | 2410 |
| abstract_inverted_index.(mid-adult | 1920 |
| abstract_inverted_index.(proband), | 1407 |
| abstract_inverted_index./ChchdDefA | 2356 |
| abstract_inverted_index.1-week-old | 1467, 1518, 1578, 2065, 2637, 3107, 3206, 3665, 3683 |
| abstract_inverted_index.1—figure | 2012, 2112, 2212, 2250 |
| abstract_inverted_index.1—source | 1523 |
| abstract_inverted_index.C17orf67), | 1726 |
| abstract_inverted_index.Dgkepsilon | 1944 |
| abstract_inverted_index.Discussion | 13 |
| abstract_inverted_index.Drosophila | 83, 705, 724, 785, 1171, 1380, 1427, 1435, 1445, 1469, 1474, 1538, 1546, 1556, 1580, 1788, 1815, 1831, 1844, 1853, 2330, 2858, 3109, 3159, 3612, 3620 |
| abstract_inverted_index.Fractional | 2633, 3677 |
| abstract_inverted_index.Late-stage | 2353 |
| abstract_inverted_index.References | 19 |
| abstract_inverted_index.Similarly, | 885 |
| abstract_inverted_index.additional | 179, 200 |
| abstract_inverted_index.aggregated | 1110 |
| abstract_inverted_index.assembly), | 1943 |
| abstract_inverted_index.associated | 609, 1039, 1749, 3830 |
| abstract_inverted_index.candidates | 1804 |
| abstract_inverted_index.cerebellar | 1330 |
| abstract_inverted_index.complexity | 673 |
| abstract_inverted_index.components | 310, 1203, 3236 |
| abstract_inverted_index.confirming | 2207, 2239 |
| abstract_inverted_index.congenital | 38, 393 |
| abstract_inverted_index.conserved, | 728 |
| abstract_inverted_index.consistent | 165 |
| abstract_inverted_index.contribute | 695, 1222 |
| abstract_inverted_index.deficiency | 2546 |
| abstract_inverted_index.determined | 629 |
| abstract_inverted_index.developing | 782 |
| abstract_inverted_index.diminished | 130, 1092, 1146, 2936, 2982, 3038, 3051, 3509, 4047, 4159 |
| abstract_inverted_index.downstream | 1756 |
| abstract_inverted_index.efficiency | 2437, 2710, 2716 |
| abstract_inverted_index.evaluation | 10, 279 |
| abstract_inverted_index.expressed. | 1493 |
| abstract_inverted_index.expression | 847, 2481, 2510, 2952, 3316 |
| abstract_inverted_index.fractional | 1514, 1974, 1993, 2085, 2186, 2237, 2443, 2599, 2754, 2765, 2821, 3340, 3714, 3732, 3987 |
| abstract_inverted_index.fragmented | 1108 |
| abstract_inverted_index.functional | 1071, 1452, 1563, 2448, 2873 |
| abstract_inverted_index.generating | 335 |
| abstract_inverted_index.highlights | 352 |
| abstract_inverted_index.homozygous | 111, 965, 1265, 1686, 1743 |
| abstract_inverted_index.horizontal | 1398 |
| abstract_inverted_index.identified | 1209, 2516 |
| abstract_inverted_index.importance | 354 |
| abstract_inverted_index.incomplete | 1767 |
| abstract_inverted_index.increased, | 2006 |
| abstract_inverted_index.individual | 1770, 3032, 3156 |
| abstract_inverted_index.machinery, | 1047 |
| abstract_inverted_index.manifested | 898, 1106 |
| abstract_inverted_index.manuscript | 363 |
| abstract_inverted_index.mechanisms | 56, 666, 920 |
| abstract_inverted_index.microscopy | 880 |
| abstract_inverted_index.molecular, | 698 |
| abstract_inverted_index.morphology | 174, 1032, 1103 |
| abstract_inverted_index.negatively | 1939 |
| abstract_inverted_index.neonatally | 524 |
| abstract_inverted_index.non-coding | 1714 |
| abstract_inverted_index.off-target | 2224 |
| abstract_inverted_index.oligogenic | 45, 193, 336, 672, 1762 |
| abstract_inverted_index.orthologs. | 1428, 1539 |
| abstract_inverted_index.paired-end | 1623 |
| abstract_inverted_index.pan-muscle | 2171, 2193, 3500 |
| abstract_inverted_index.pathogenic | 55, 665 |
| abstract_inverted_index.penetrance | 1768 |
| abstract_inverted_index.phenotypes | 346, 466, 1113, 1453, 1564 |
| abstract_inverted_index.polymerase | 227, 1183 |
| abstract_inverted_index.postulated | 800, 1272, 1761 |
| abstract_inverted_index.previously | 1347, 2019 |
| abstract_inverted_index.procedure, | 521 |
| abstract_inverted_index.procedures | 550 |
| abstract_inverted_index.processes, | 853 |
| abstract_inverted_index.regulating | 1940 |
| abstract_inverted_index.respective | 1830 |
| abstract_inverted_index.sarcomeric | 133, 1093, 2865, 2896, 3023, 3033, 3085, 3094, 3211, 3240, 3297, 3381, 3413, 3474 |
| abstract_inverted_index.sensitized | 213 |
| abstract_inverted_index.sequencing | 66, 330, 948, 1587 |
| abstract_inverted_index.shortening | 1515, 2187, 2444, 2600, 2634, 2755, 2766, 3678, 3733, 3988 |
| abstract_inverted_index.standards; | 1629 |
| abstract_inverted_index.strategy). | 2725 |
| abstract_inverted_index.structural | 1285 |
| abstract_inverted_index.sufficient | 2468 |
| abstract_inverted_index.suggesting | 243 |
| abstract_inverted_index.supplement | 1365, 2013, 2113, 2213, 2251, 3067, 3326, 3352, 3390, 3723, 3756, 3785, 3818, 4016, 4036, 4058, 4086, 4090, 4103, 4141, 4165, 4190, 4216, 4227 |
| abstract_inverted_index.throughout | 1899, 2431, 2830 |
| abstract_inverted_index.transport. | 860 |
| abstract_inverted_index.treatments | 2815 |
| abstract_inverted_index.two-tailed | 2681, 3223 |
| abstract_inverted_index.unaffected | 1746, 3005 |
| abstract_inverted_index.underlying | 703, 921, 931 |
| abstract_inverted_index.understand | 691, 2265 |
| abstract_inverted_index.ventricle, | 559 |
| abstract_inverted_index.(ChchdDefA) | 2548 |
| abstract_inverted_index.1000–2000 | 399 |
| abstract_inverted_index.CB-specific | 2504 |
| abstract_inverted_index.Drosophila) | 1019 |
| abstract_inverted_index.Drosophila, | 981, 3926 |
| abstract_inverted_index.Hypoplastic | 30, 379 |
| abstract_inverted_index.activator), | 1185 |
| abstract_inverted_index.cardiogenic | 734, 2336 |
| abstract_inverted_index.chromosomal | 1601 |
| abstract_inverted_index.circulation | 533 |
| abstract_inverted_index.combination | 221, 3709, 3742, 3770, 3799 |
| abstract_inverted_index.comparative | 1595 |
| abstract_inverted_index.complex’s | 169, 1065 |
| abstract_inverted_index.compromised | 127, 4255 |
| abstract_inverted_index.contractile | 837, 2881, 2891, 3052, 3451 |
| abstract_inverted_index.development | 1069 |
| abstract_inverted_index.drastically | 126, 3765 |
| abstract_inverted_index.duplication | 1604 |
| abstract_inverted_index.dysfunction | 1352, 2010, 2094 |
| abstract_inverted_index.elucidation | 253 |
| abstract_inverted_index.epigenetic, | 417 |
| abstract_inverted_index.established | 1814 |
| abstract_inverted_index.experiments | 288 |
| abstract_inverted_index.expression, | 2027, 3259 |
| abstract_inverted_index.genetically | 1176, 3905 |
| abstract_inverted_index.homeostasis | 1072, 1207 |
| abstract_inverted_index.identifying | 325 |
| abstract_inverted_index.independent | 2075 |
| abstract_inverted_index.information | 27 |
| abstract_inverted_index.inheritance | 1270 |
| abstract_inverted_index.interaction | 1168, 3720, 3751, 3782, 3810, 3923, 4031, 4136, 4220 |
| abstract_inverted_index.investigate | 2690 |
| abstract_inverted_index.involvement | 246 |
| abstract_inverted_index.maintaining | 172, 1215, 2258, 2851 |
| abstract_inverted_index.oxygen-poor | 536 |
| abstract_inverted_index.palliation. | 1231, 1317 |
| abstract_inverted_index.pericardial | 1486, 1906, 2158 |
| abstract_inverted_index.phenotypes. | 2352 |
| abstract_inverted_index.pluripotent | 1151 |
| abstract_inverted_index.potentially | 262 |
| abstract_inverted_index.preparation | 2123 |
| abstract_inverted_index.prioritized | 103, 201, 956 |
| abstract_inverted_index.production, | 835 |
| abstract_inverted_index.production. | 1035, 1083 |
| abstract_inverted_index.projection) | 2501 |
| abstract_inverted_index.requirement | 1790, 2693 |
| abstract_inverted_index.scaffolding | 237, 1196 |
| abstract_inverted_index.semi-intact | 2120 |
| abstract_inverted_index.shortening, | 1975, 1994, 2086, 2238, 2822, 3341, 3715 |
| abstract_inverted_index.significant | 1806, 2596, 3718, 3780, 3981, 4241 |
| abstract_inverted_index.single-cell | 2316, 2505 |
| abstract_inverted_index.structures. | 1111 |
| abstract_inverted_index.substantial | 2818, 3365 |
| abstract_inverted_index.synergistic | 240 |
| abstract_inverted_index.temperature | 2629 |
| abstract_inverted_index.three-stage | 519 |
| abstract_inverted_index.ventricular | 554, 828, 1310 |
| abstract_inverted_index.well-suited | 707 |
| abstract_inverted_index.End-systolic | 1509 |
| abstract_inverted_index.Hand4.2-Gal4 | 1449, 1490, 1560, 1880, 1932, 2066, 2228, 2726, 2951, 3112, 3162 |
| abstract_inverted_index.Introduction | 11, 378 |
| abstract_inverted_index.Mitochondria | 797 |
| abstract_inverted_index.availability | 18 |
| abstract_inverted_index.cardioblasts | 2331 |
| abstract_inverted_index.complexities | 53 |
| abstract_inverted_index.conclusively | 628 |
| abstract_inverted_index.demonstrated | 349, 1672 |
| abstract_inverted_index.development, | 2311, 2698, 2844 |
| abstract_inverted_index.establishing | 1213, 1794 |
| abstract_inverted_index.experimental | 925, 2724 |
| abstract_inverted_index.experiments. | 2631, 3924 |
| abstract_inverted_index.family-based | 1165 |
| abstract_inverted_index.functionally | 79, 683 |
| abstract_inverted_index.highlighting | 1475 |
| abstract_inverted_index.hypothesized | 2029, 3261, 3447 |
| abstract_inverted_index.iPSC-derived | 293 |
| abstract_inverted_index.individually | 1442, 1553 |
| abstract_inverted_index.interactions | 209, 260, 4243 |
| abstract_inverted_index.investigated | 968 |
| abstract_inverted_index.measurements | 2635 |
| abstract_inverted_index.mitochondria | 934, 3444, 3484, 3610 |
| abstract_inverted_index.organization | 998 |
| abstract_inverted_index.pan-neuronal | 2176 |
| abstract_inverted_index.post-mitotic | 1892 |
| abstract_inverted_index.requirements | 1807, 2254, 2271 |
| abstract_inverted_index.specifically | 1136 |
| abstract_inverted_index.transmission | 878 |
| abstract_inverted_index.untranslated | 1722 |
| abstract_inverted_index.(β-Spectrin, | 236 |
| abstract_inverted_index.Bioinformatic | 86 |
| abstract_inverted_index.Chchd3/6RNAiA | 2216, 2730 |
| abstract_inverted_index.Chchd3/6RNAiB | 2218 |
| abstract_inverted_index.End-Diastolic | 1505 |
| abstract_inverted_index.Gal4-mediated | 2706 |
| abstract_inverted_index.Hypothesizing | 191 |
| abstract_inverted_index.approximately | 758 |
| abstract_inverted_index.approximation | 2499 |
| abstract_inverted_index.arrhythmicity | 1977 |
| abstract_inverted_index.bioinformatic | 927 |
| abstract_inverted_index.characterized | 1258 |
| abstract_inverted_index.consanguinity | 1276 |
| abstract_inverted_index.contractility | 1087, 2017, 2034, 2090, 2733, 3803, 3982, 4025, 4078, 4179, 4201, 4234 |
| abstract_inverted_index.corresponding | 1426, 1537 |
| abstract_inverted_index.developmental | 788, 1326, 2780 |
| abstract_inverted_index.downregulated | 846 |
| abstract_inverted_index.eight-subunit | 1013 |
| abstract_inverted_index.environmental | 419 |
| abstract_inverted_index.establishment | 2418 |
| abstract_inverted_index.hybridization | 1597 |
| abstract_inverted_index.mitochondrial | 119, 142, 832, 852, 869, 874, 896, 905, 919, 1024, 1055, 1102, 1128, 1206, 1351, 2025, 2039, 3274, 3466, 3471, 3544, 3570, 3688, 3853, 3876, 3896, 3913, 3933, 3946, 4245, 4260 |
| abstract_inverted_index.morphological | 701 |
| abstract_inverted_index.pathogenesis, | 923 |
| abstract_inverted_index.pathogenesis. | 808 |
| abstract_inverted_index.proliferation | 1147 |
| abstract_inverted_index.significantly | 1145, 1456, 1567 |
| abstract_inverted_index.similarities, | 789 |
| abstract_inverted_index.specification | 2381 |
| abstract_inverted_index.transcription | 735, 2365, 2555 |
| abstract_inverted_index.underpinnings | 602, 711 |
| abstract_inverted_index.understanding | 49, 272, 915 |
| abstract_inverted_index.(Mitochondrial | 1951 |
| abstract_inverted_index.(iPSC)-derived | 1154 |
| abstract_inverted_index.(mitochondrial | 993 |
| abstract_inverted_index.>Chchd3/6RNAiA | 2426, 2591, 2615, 3872 |
| abstract_inverted_index.Chchd3/6RNAiC1 | 2746, 3744 |
| abstract_inverted_index.Interestingly, | 2813 |
| abstract_inverted_index.Nkx2-5/tinman) | 741 |
| abstract_inverted_index.Prioritization | 1372 |
| abstract_inverted_index.Three-week-old | 1919 |
| abstract_inverted_index.bioinformatics | 1589 |
| abstract_inverted_index.cardiogenesis) | 2530 |
| abstract_inverted_index.cardiomyocytes | 294, 816, 1156, 1903, 2884, 2957 |
| abstract_inverted_index.concentration, | 833 |
| abstract_inverted_index.consanguineous | 101, 957, 1393 |
| abstract_inverted_index.contractility, | 129, 3767 |
| abstract_inverted_index.contractility. | 319, 1999 |
| abstract_inverted_index.evolutionarily | 727 |
| abstract_inverted_index.fission-fusion | 143, 3571, 3597, 3689, 3854, 3877, 3899, 3914, 4246 |
| abstract_inverted_index.heart-specific | 1894, 2315, 3077 |
| abstract_inverted_index.non-autonomous | 2144 |
| abstract_inverted_index.patient-parent | 71 |
| abstract_inverted_index.phenotypically | 1257 |
| abstract_inverted_index.proband-parent | 943 |
| abstract_inverted_index.reconstructive | 492 |
| abstract_inverted_index.transcriptomic | 2317 |
| abstract_inverted_index.(CHCHD6)(Marian | 1729 |
| abstract_inverted_index.****p≤0.0001, | 2683 |
| abstract_inverted_index.Tomita-Mitchell | 650 |
| abstract_inverted_index.characteristics | 440 |
| abstract_inverted_index.disease-causing | 763 |
| abstract_inverted_index.pathophysiology | 471 |
| abstract_inverted_index.patient-derived | 865 |
| abstract_inverted_index.transcriptomics | 2506 |
| abstract_inverted_index.Kozjak-Pavlovic, | 1061 |
| abstract_inverted_index.cardiac-specific | 115, 982, 1089, 1819, 2137 |
| abstract_inverted_index.echocardiograms. | 1300 |
| abstract_inverted_index.echocardiography | 1260 |
| abstract_inverted_index.or>Chchd3/6RNAiA | 2572 |
| abstract_inverted_index.transplantation, | 497 |
| abstract_inverted_index.ventricular-like | 1155 |
| abstract_inverted_index.Martínez-Morentin | 2045 |
| abstract_inverted_index.trans-heterozygous | 2357 |
| abstract_inverted_index.ventricle-dependent | 531 |
| abstract_inverted_index.disease-contributing | 263 |
| abstract_inverted_index.Hand4.2-Gal4>Chchd3/6 | 1520, 2942 |
| abstract_inverted_index.temperature-dependence | 2704 |
| abstract_inverted_index.cardiomyocyte-autonomous | 2209 |
| abstract_inverted_index.mesoderm/muscle-specific | 2364 |
| abstract_inverted_index.Hand4.2-Gal4>Chchd3/6RNAiC1 | 2782 |
| abstract_inverted_index.elife-83385-fig1-data1-v2.xlsx | 1584 |
| abstract_inverted_index.https://doi.org/10.7554/eLife.83385.sa0 | 372 |
| abstract_inverted_index.(coiled-coil-helix-coiled-coil-helix-domain-containing | 987 |
| abstract_inverted_index.https://cdn.elifesciences.org/articles/83385/elife-83385-fig1-data1-v2.xlsx | 1582 |
| cited_by_percentile_year | |
| countries_distinct_count | 1 |
| institutions_distinct_count | 16 |
| sustainable_development_goals[0].id | https://metadata.un.org/sdg/3 |
| sustainable_development_goals[0].score | 0.47999998927116394 |
| sustainable_development_goals[0].display_name | Good health and well-being |
| citation_normalized_percentile |