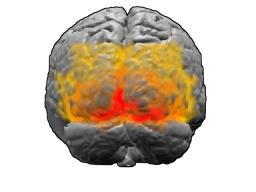
EEL human visual cortex 445 genes RNA spatial data Article Swipe
Lars E. Borm
,
Alejandro Mossi Albiach
·
 YOU?
·
· 2022
· Open Access
·
· DOI: https://doi.org/10.6084/m9.figshare.20324865.v2
YOU?
·
· 2022
· Open Access
·
· DOI: https://doi.org/10.6084/m9.figshare.20324865.v2
 YOU?
·
· 2022
· Open Access
·
· DOI: https://doi.org/10.6084/m9.figshare.20324865.v2
YOU?
·
· 2022
· Open Access
·
· DOI: https://doi.org/10.6084/m9.figshare.20324865.v2
Raw RNA locations of a visual cortex human brain section produced by EEL FISH for 445 genes. RNA files are in the .parquet format which can be opened with FISHscale (https://github.com/linnarsson-lab/FISHscale) or any other parquet file reader (https://arrow.apache.org/docs/index.html) RNA .parquet files Position and gene label for all RNA molecules. "c_px_microscope_stitched" contains X coordinates. "r_px_microscope_stitched" contians Y coordinates. The unit are pixels with a size of 0.18 micrometer. Multiply by 0.18 to get um scale. "hamming_distance" is the hamming distance divided by the barcode length (16). The closer to 0 the better the barcode. Gene colors .pkl file Pickled Python dictionary with gene colors used in the paper.
Related Topics
Concepts
Metadata
- Type
- dataset
- Language
- en
- Landing Page
- https://doi.org/10.6084/m9.figshare.20324865.v2
- OA Status
- gold
- Related Works
- 10
- OpenAlex ID
- https://openalex.org/W4394176081
All OpenAlex metadata
Raw OpenAlex JSON
- OpenAlex ID
-
https://openalex.org/W4394176081Canonical identifier for this work in OpenAlex
- DOI
-
https://doi.org/10.6084/m9.figshare.20324865.v2Digital Object Identifier
- Title
-
EEL human visual cortex 445 genes RNA spatial dataWork title
- Type
-
datasetOpenAlex work type
- Language
-
enPrimary language
- Publication year
-
2022Year of publication
- Publication date
-
2022-01-01Full publication date if available
- Authors
-
Lars E. Borm, Alejandro Mossi AlbiachList of authors in order
- Landing page
-
https://doi.org/10.6084/m9.figshare.20324865.v2Publisher landing page
- Open access
-
YesWhether a free full text is available
- OA status
-
goldOpen access status per OpenAlex
- OA URL
-
https://doi.org/10.6084/m9.figshare.20324865.v2Direct OA link when available
- Concepts
-
Cortex (anatomy), Gene, Visual cortex, Computational biology, RNA, Biology, Cartography, Computer science, Neuroscience, Geography, GeneticsTop concepts (fields/topics) attached by OpenAlex
- Cited by
-
0Total citation count in OpenAlex
- Related works (count)
-
10Other works algorithmically related by OpenAlex
Full payload
| id | https://openalex.org/W4394176081 |
|---|---|
| doi | https://doi.org/10.6084/m9.figshare.20324865.v2 |
| ids.doi | https://doi.org/10.6084/m9.figshare.20324865.v2 |
| ids.openalex | https://openalex.org/W4394176081 |
| fwci | 0.0 |
| type | dataset |
| title | EEL human visual cortex 445 genes RNA spatial data |
| biblio.issue | |
| biblio.volume | |
| biblio.last_page | |
| biblio.first_page | |
| topics[0].id | https://openalex.org/T11289 |
| topics[0].field.id | https://openalex.org/fields/13 |
| topics[0].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[0].score | 0.7616000175476074 |
| topics[0].domain.id | https://openalex.org/domains/1 |
| topics[0].domain.display_name | Life Sciences |
| topics[0].subfield.id | https://openalex.org/subfields/1312 |
| topics[0].subfield.display_name | Molecular Biology |
| topics[0].display_name | Single-cell and spatial transcriptomics |
| topics[1].id | https://openalex.org/T12859 |
| topics[1].field.id | https://openalex.org/fields/13 |
| topics[1].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[1].score | 0.6503000259399414 |
| topics[1].domain.id | https://openalex.org/domains/1 |
| topics[1].domain.display_name | Life Sciences |
| topics[1].subfield.id | https://openalex.org/subfields/1304 |
| topics[1].subfield.display_name | Biophysics |
| topics[1].display_name | Cell Image Analysis Techniques |
| is_xpac | False |
| apc_list | |
| apc_paid | |
| concepts[0].id | https://openalex.org/C2777348757 |
| concepts[0].level | 2 |
| concepts[0].score | 0.5067897439002991 |
| concepts[0].wikidata | https://www.wikidata.org/wiki/Q2346577 |
| concepts[0].display_name | Cortex (anatomy) |
| concepts[1].id | https://openalex.org/C104317684 |
| concepts[1].level | 2 |
| concepts[1].score | 0.4884432852268219 |
| concepts[1].wikidata | https://www.wikidata.org/wiki/Q7187 |
| concepts[1].display_name | Gene |
| concepts[2].id | https://openalex.org/C2779345533 |
| concepts[2].level | 2 |
| concepts[2].score | 0.46564897894859314 |
| concepts[2].wikidata | https://www.wikidata.org/wiki/Q75785 |
| concepts[2].display_name | Visual cortex |
| concepts[3].id | https://openalex.org/C70721500 |
| concepts[3].level | 1 |
| concepts[3].score | 0.45401445031166077 |
| concepts[3].wikidata | https://www.wikidata.org/wiki/Q177005 |
| concepts[3].display_name | Computational biology |
| concepts[4].id | https://openalex.org/C67705224 |
| concepts[4].level | 3 |
| concepts[4].score | 0.42550450563430786 |
| concepts[4].wikidata | https://www.wikidata.org/wiki/Q11053 |
| concepts[4].display_name | RNA |
| concepts[5].id | https://openalex.org/C86803240 |
| concepts[5].level | 0 |
| concepts[5].score | 0.4163048267364502 |
| concepts[5].wikidata | https://www.wikidata.org/wiki/Q420 |
| concepts[5].display_name | Biology |
| concepts[6].id | https://openalex.org/C58640448 |
| concepts[6].level | 1 |
| concepts[6].score | 0.34749042987823486 |
| concepts[6].wikidata | https://www.wikidata.org/wiki/Q42515 |
| concepts[6].display_name | Cartography |
| concepts[7].id | https://openalex.org/C41008148 |
| concepts[7].level | 0 |
| concepts[7].score | 0.34705159068107605 |
| concepts[7].wikidata | https://www.wikidata.org/wiki/Q21198 |
| concepts[7].display_name | Computer science |
| concepts[8].id | https://openalex.org/C169760540 |
| concepts[8].level | 1 |
| concepts[8].score | 0.33757299184799194 |
| concepts[8].wikidata | https://www.wikidata.org/wiki/Q207011 |
| concepts[8].display_name | Neuroscience |
| concepts[9].id | https://openalex.org/C205649164 |
| concepts[9].level | 0 |
| concepts[9].score | 0.2601160705089569 |
| concepts[9].wikidata | https://www.wikidata.org/wiki/Q1071 |
| concepts[9].display_name | Geography |
| concepts[10].id | https://openalex.org/C54355233 |
| concepts[10].level | 1 |
| concepts[10].score | 0.2249164879322052 |
| concepts[10].wikidata | https://www.wikidata.org/wiki/Q7162 |
| concepts[10].display_name | Genetics |
| keywords[0].id | https://openalex.org/keywords/cortex |
| keywords[0].score | 0.5067897439002991 |
| keywords[0].display_name | Cortex (anatomy) |
| keywords[1].id | https://openalex.org/keywords/gene |
| keywords[1].score | 0.4884432852268219 |
| keywords[1].display_name | Gene |
| keywords[2].id | https://openalex.org/keywords/visual-cortex |
| keywords[2].score | 0.46564897894859314 |
| keywords[2].display_name | Visual cortex |
| keywords[3].id | https://openalex.org/keywords/computational-biology |
| keywords[3].score | 0.45401445031166077 |
| keywords[3].display_name | Computational biology |
| keywords[4].id | https://openalex.org/keywords/rna |
| keywords[4].score | 0.42550450563430786 |
| keywords[4].display_name | RNA |
| keywords[5].id | https://openalex.org/keywords/biology |
| keywords[5].score | 0.4163048267364502 |
| keywords[5].display_name | Biology |
| keywords[6].id | https://openalex.org/keywords/cartography |
| keywords[6].score | 0.34749042987823486 |
| keywords[6].display_name | Cartography |
| keywords[7].id | https://openalex.org/keywords/computer-science |
| keywords[7].score | 0.34705159068107605 |
| keywords[7].display_name | Computer science |
| keywords[8].id | https://openalex.org/keywords/neuroscience |
| keywords[8].score | 0.33757299184799194 |
| keywords[8].display_name | Neuroscience |
| keywords[9].id | https://openalex.org/keywords/geography |
| keywords[9].score | 0.2601160705089569 |
| keywords[9].display_name | Geography |
| keywords[10].id | https://openalex.org/keywords/genetics |
| keywords[10].score | 0.2249164879322052 |
| keywords[10].display_name | Genetics |
| language | en |
| locations[0].id | doi:10.6084/m9.figshare.20324865.v2 |
| locations[0].is_oa | True |
| locations[0].source | |
| locations[0].license | cc-by |
| locations[0].pdf_url | |
| locations[0].version | |
| locations[0].raw_type | dataset |
| locations[0].license_id | https://openalex.org/licenses/cc-by |
| locations[0].is_accepted | False |
| locations[0].is_published | |
| locations[0].raw_source_name | |
| locations[0].landing_page_url | https://doi.org/10.6084/m9.figshare.20324865.v2 |
| indexed_in | datacite |
| authorships[0].author.id | https://openalex.org/A5058580972 |
| authorships[0].author.orcid | https://orcid.org/0000-0002-1764-6057 |
| authorships[0].author.display_name | Lars E. Borm |
| authorships[0].author_position | first |
| authorships[0].raw_author_name | Lars Borm |
| authorships[0].is_corresponding | False |
| authorships[1].author.id | https://openalex.org/A5081753875 |
| authorships[1].author.orcid | https://orcid.org/0000-0001-7009-4901 |
| authorships[1].author.display_name | Alejandro Mossi Albiach |
| authorships[1].author_position | last |
| authorships[1].raw_author_name | Alejandro Mossi Albiach |
| authorships[1].is_corresponding | False |
| has_content.pdf | False |
| has_content.grobid_xml | False |
| is_paratext | False |
| open_access.is_oa | True |
| open_access.oa_url | https://doi.org/10.6084/m9.figshare.20324865.v2 |
| open_access.oa_status | gold |
| open_access.any_repository_has_fulltext | False |
| created_date | 2025-10-10T00:00:00 |
| display_name | EEL human visual cortex 445 genes RNA spatial data |
| has_fulltext | False |
| is_retracted | False |
| updated_date | 2025-11-06T06:51:31.235846 |
| primary_topic.id | https://openalex.org/T11289 |
| primary_topic.field.id | https://openalex.org/fields/13 |
| primary_topic.field.display_name | Biochemistry, Genetics and Molecular Biology |
| primary_topic.score | 0.7616000175476074 |
| primary_topic.domain.id | https://openalex.org/domains/1 |
| primary_topic.domain.display_name | Life Sciences |
| primary_topic.subfield.id | https://openalex.org/subfields/1312 |
| primary_topic.subfield.display_name | Molecular Biology |
| primary_topic.display_name | Single-cell and spatial transcriptomics |
| related_works | https://openalex.org/W2147993839, https://openalex.org/W2034574984, https://openalex.org/W1993532945, https://openalex.org/W1994056074, https://openalex.org/W1995132808, https://openalex.org/W2015845327, https://openalex.org/W2412669475, https://openalex.org/W1986370492, https://openalex.org/W2160579003, https://openalex.org/W1485633775 |
| cited_by_count | 0 |
| locations_count | 1 |
| best_oa_location.id | doi:10.6084/m9.figshare.20324865.v2 |
| best_oa_location.is_oa | True |
| best_oa_location.source | |
| best_oa_location.license | cc-by |
| best_oa_location.pdf_url | |
| best_oa_location.version | |
| best_oa_location.raw_type | dataset |
| best_oa_location.license_id | https://openalex.org/licenses/cc-by |
| best_oa_location.is_accepted | False |
| best_oa_location.is_published | False |
| best_oa_location.raw_source_name | |
| best_oa_location.landing_page_url | https://doi.org/10.6084/m9.figshare.20324865.v2 |
| primary_location.id | doi:10.6084/m9.figshare.20324865.v2 |
| primary_location.is_oa | True |
| primary_location.source | |
| primary_location.license | cc-by |
| primary_location.pdf_url | |
| primary_location.version | |
| primary_location.raw_type | dataset |
| primary_location.license_id | https://openalex.org/licenses/cc-by |
| primary_location.is_accepted | False |
| primary_location.is_published | False |
| primary_location.raw_source_name | |
| primary_location.landing_page_url | https://doi.org/10.6084/m9.figshare.20324865.v2 |
| publication_date | 2022-01-01 |
| publication_year | 2022 |
| referenced_works_count | 0 |
| abstract_inverted_index.0 | 91 |
| abstract_inverted_index.X | 53 |
| abstract_inverted_index.Y | 57 |
| abstract_inverted_index.a | 4, 64 |
| abstract_inverted_index.be | 27 |
| abstract_inverted_index.by | 11, 70, 83 |
| abstract_inverted_index.in | 21, 108 |
| abstract_inverted_index.is | 78 |
| abstract_inverted_index.of | 3, 66 |
| abstract_inverted_index.or | 32 |
| abstract_inverted_index.to | 72, 90 |
| abstract_inverted_index.um | 74 |
| abstract_inverted_index.445 | 15 |
| abstract_inverted_index.EEL | 12 |
| abstract_inverted_index.RNA | 1, 18, 49 |
| abstract_inverted_index.The | 59, 88 |
| abstract_inverted_index.all | 48 |
| abstract_inverted_index.and | 44 |
| abstract_inverted_index.any | 33 |
| abstract_inverted_index.are | 20, 61 |
| abstract_inverted_index.can | 26 |
| abstract_inverted_index.for | 14, 47 |
| abstract_inverted_index.get | 73 |
| abstract_inverted_index.the | 22, 79, 84, 92, 94, 109 |
| abstract_inverted_index..pkl | 99 |
| abstract_inverted_index.0.18 | 67, 71 |
| abstract_inverted_index.<br> | 17, 39, 76, 96 |
| abstract_inverted_index.FISH | 13 |
| abstract_inverted_index.file | 36 |
| abstract_inverted_index.gene | 45, 105 |
| abstract_inverted_index.size | 65 |
| abstract_inverted_index.unit | 60 |
| abstract_inverted_index.used | 107 |
| abstract_inverted_index.with | 29, 63, 104 |
| abstract_inverted_index.(16). | 87 |
| abstract_inverted_index.brain | 8 |
| abstract_inverted_index.files | 19 |
| abstract_inverted_index.human | 7 |
| abstract_inverted_index.label | 46 |
| abstract_inverted_index.other | 34 |
| abstract_inverted_index.which | 25 |
| abstract_inverted_index.Python | 102 |
| abstract_inverted_index.better | 93 |
| abstract_inverted_index.closer | 89 |
| abstract_inverted_index.colors | 98, 106 |
| abstract_inverted_index.cortex | 6 |
| abstract_inverted_index.format | 24 |
| abstract_inverted_index.length | 86 |
| abstract_inverted_index.opened | 28 |
| abstract_inverted_index.paper. | 110 |
| abstract_inverted_index.pixels | 62 |
| abstract_inverted_index.reader | 37 |
| abstract_inverted_index.scale. | 75 |
| abstract_inverted_index.visual | 5 |
| abstract_inverted_index.Pickled | 101 |
| abstract_inverted_index.barcode | 85 |
| abstract_inverted_index.divided | 82 |
| abstract_inverted_index.hamming | 80 |
| abstract_inverted_index.parquet | 35 |
| abstract_inverted_index.section | 9 |
| abstract_inverted_index..parquet | 23, 41 |
| abstract_inverted_index.Multiply | 69 |
| abstract_inverted_index.Position | 43 |
| abstract_inverted_index.barcode. | 95 |
| abstract_inverted_index.contains | 52 |
| abstract_inverted_index.contians | 56 |
| abstract_inverted_index.distance | 81 |
| abstract_inverted_index.produced | 10 |
| abstract_inverted_index.FISHscale | 30 |
| abstract_inverted_index.locations | 2 |
| abstract_inverted_index.dictionary | 103 |
| abstract_inverted_index.molecules. | 50 |
| abstract_inverted_index.<strong>RNA | 40 |
| abstract_inverted_index.<strong>Raw | 0 |
| abstract_inverted_index.micrometer. | 68 |
| abstract_inverted_index.<strong>Gene | 97 |
| abstract_inverted_index.coordinates. | 54, 58 |
| abstract_inverted_index.file</strong> | 100 |
| abstract_inverted_index.files</strong> | 42 |
| abstract_inverted_index.genes.</strong> | 16 |
| abstract_inverted_index."hamming_distance" | 77 |
| abstract_inverted_index."c_px_microscope_stitched" | 51 |
| abstract_inverted_index."r_px_microscope_stitched" | 55 |
| abstract_inverted_index.(https://arrow.apache.org/docs/index.html) | 38 |
| abstract_inverted_index.(https://github.com/linnarsson-lab/FISHscale) | 31 |
| cited_by_percentile_year | |
| countries_distinct_count | 0 |
| institutions_distinct_count | 2 |
| citation_normalized_percentile |