eLife Assessment: Near-perfect precise on-target editing of human hematopoietic stem and progenitor cells Article Swipe
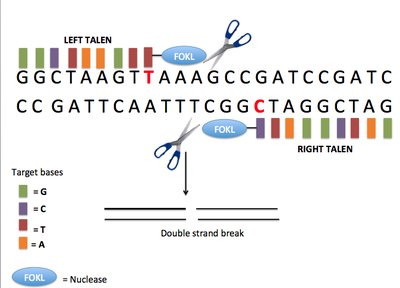
Precision gene editing in primary hematopoietic stem and progenitor cells (HSPCs) would facilitate both curative treatments for monogenic disorders as well as disease modelling. Precise efficiencies even with the CRISPR/Cas system, however, remain limited. Through an optimization of guide RNA delivery, donor design, and additives, we have now obtained mean precise editing efficiencies >90% on primary cord blood HSCPs with minimal toxicity and without observed off-target editing. The main protocol modifications needed to achieve such high efficiencies were the addition of the DNA-PK inhibitor AZD7648, and the inclusion of spacer-breaking silent mutations in the donor in addition to mutations disrupting the PAM sequence. Critically, editing was even across the progenitor hierarchy, and did not substantially distort the hierarchy or affect lineage outputs in colony-forming cell assays. As modelling of many diseases requires heterozygosity, we also demonstrated that the overall editing and zygosity can be tuned by adding in defined mixtures of mutant and wild-type donor. With these optimizations, editing at near-perfect efficiency can now be accomplished directly in human HSPCs. This will open new avenues in both therapeutic strategies and disease modelling.
Related Topics
- Type
- peer-review
- Language
- en
- Landing Page
- https://doi.org/10.7554/elife.91288.1.sa2
- OA Status
- gold
- Related Works
- 10
- OpenAlex ID
- https://openalex.org/W4387360728
Raw OpenAlex JSON
- OpenAlex ID
-
https://openalex.org/W4387360728Canonical identifier for this work in OpenAlex
- DOI
-
https://doi.org/10.7554/elife.91288.1.sa2Digital Object Identifier
- Title
-
eLife Assessment: Near-perfect precise on-target editing of human hematopoietic stem and progenitor cellsWork title
- Type
-
peer-reviewOpenAlex work type
- Language
-
enPrimary language
- Publication year
-
2023Year of publication
- Publication date
-
2023-10-05Full publication date if available
- Authors
-
Jian XuList of authors in order
- Landing page
-
https://doi.org/10.7554/elife.91288.1.sa2Publisher landing page
- Open access
-
YesWhether a free full text is available
- OA status
-
goldOpen access status per OpenAlex
- OA URL
-
https://doi.org/10.7554/elife.91288.1.sa2Direct OA link when available
- Concepts
-
Genome editing, Progenitor cell, Stem cell, Computational biology, Biology, Haematopoiesis, CRISPR, Cord blood, Primary cell, Genetics, Computer science, GeneTop concepts (fields/topics) attached by OpenAlex
- Cited by
-
0Total citation count in OpenAlex
- Related works (count)
-
10Other works algorithmically related by OpenAlex
Full payload
| id | https://openalex.org/W4387360728 |
|---|---|
| doi | https://doi.org/10.7554/elife.91288.1.sa2 |
| ids.doi | https://doi.org/10.7554/elife.91288.1.sa2 |
| ids.openalex | https://openalex.org/W4387360728 |
| fwci | 0.0 |
| type | peer-review |
| title | eLife Assessment: Near-perfect precise on-target editing of human hematopoietic stem and progenitor cells |
| biblio.issue | |
| biblio.volume | |
| biblio.last_page | |
| biblio.first_page | |
| topics[0].id | https://openalex.org/T10878 |
| topics[0].field.id | https://openalex.org/fields/13 |
| topics[0].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[0].score | 1.0 |
| topics[0].domain.id | https://openalex.org/domains/1 |
| topics[0].domain.display_name | Life Sciences |
| topics[0].subfield.id | https://openalex.org/subfields/1312 |
| topics[0].subfield.display_name | Molecular Biology |
| topics[0].display_name | CRISPR and Genetic Engineering |
| topics[1].id | https://openalex.org/T10613 |
| topics[1].field.id | https://openalex.org/fields/13 |
| topics[1].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[1].score | 0.996999979019165 |
| topics[1].domain.id | https://openalex.org/domains/1 |
| topics[1].domain.display_name | Life Sciences |
| topics[1].subfield.id | https://openalex.org/subfields/1311 |
| topics[1].subfield.display_name | Genetics |
| topics[1].display_name | Virus-based gene therapy research |
| topics[2].id | https://openalex.org/T10981 |
| topics[2].field.id | https://openalex.org/fields/27 |
| topics[2].field.display_name | Medicine |
| topics[2].score | 0.9897000193595886 |
| topics[2].domain.id | https://openalex.org/domains/4 |
| topics[2].domain.display_name | Health Sciences |
| topics[2].subfield.id | https://openalex.org/subfields/2713 |
| topics[2].subfield.display_name | Epidemiology |
| topics[2].display_name | Cytomegalovirus and herpesvirus research |
| is_xpac | False |
| apc_list | |
| apc_paid | |
| concepts[0].id | https://openalex.org/C144501496 |
| concepts[0].level | 4 |
| concepts[0].score | 0.8775659799575806 |
| concepts[0].wikidata | https://www.wikidata.org/wiki/Q5533489 |
| concepts[0].display_name | Genome editing |
| concepts[1].id | https://openalex.org/C201750760 |
| concepts[1].level | 3 |
| concepts[1].score | 0.667119026184082 |
| concepts[1].wikidata | https://www.wikidata.org/wiki/Q1896218 |
| concepts[1].display_name | Progenitor cell |
| concepts[2].id | https://openalex.org/C28328180 |
| concepts[2].level | 2 |
| concepts[2].score | 0.5920475721359253 |
| concepts[2].wikidata | https://www.wikidata.org/wiki/Q48196 |
| concepts[2].display_name | Stem cell |
| concepts[3].id | https://openalex.org/C70721500 |
| concepts[3].level | 1 |
| concepts[3].score | 0.5532165765762329 |
| concepts[3].wikidata | https://www.wikidata.org/wiki/Q177005 |
| concepts[3].display_name | Computational biology |
| concepts[4].id | https://openalex.org/C86803240 |
| concepts[4].level | 0 |
| concepts[4].score | 0.5474513173103333 |
| concepts[4].wikidata | https://www.wikidata.org/wiki/Q420 |
| concepts[4].display_name | Biology |
| concepts[5].id | https://openalex.org/C109159458 |
| concepts[5].level | 3 |
| concepts[5].score | 0.5007486343383789 |
| concepts[5].wikidata | https://www.wikidata.org/wiki/Q919283 |
| concepts[5].display_name | Haematopoiesis |
| concepts[6].id | https://openalex.org/C98108389 |
| concepts[6].level | 3 |
| concepts[6].score | 0.4867730140686035 |
| concepts[6].wikidata | https://www.wikidata.org/wiki/Q412563 |
| concepts[6].display_name | CRISPR |
| concepts[7].id | https://openalex.org/C2780914630 |
| concepts[7].level | 2 |
| concepts[7].score | 0.4371558129787445 |
| concepts[7].wikidata | https://www.wikidata.org/wiki/Q914952 |
| concepts[7].display_name | Cord blood |
| concepts[8].id | https://openalex.org/C147905033 |
| concepts[8].level | 3 |
| concepts[8].score | 0.41157066822052 |
| concepts[8].wikidata | https://www.wikidata.org/wiki/Q1378887 |
| concepts[8].display_name | Primary cell |
| concepts[9].id | https://openalex.org/C54355233 |
| concepts[9].level | 1 |
| concepts[9].score | 0.3713715076446533 |
| concepts[9].wikidata | https://www.wikidata.org/wiki/Q7162 |
| concepts[9].display_name | Genetics |
| concepts[10].id | https://openalex.org/C41008148 |
| concepts[10].level | 0 |
| concepts[10].score | 0.3530680537223816 |
| concepts[10].wikidata | https://www.wikidata.org/wiki/Q21198 |
| concepts[10].display_name | Computer science |
| concepts[11].id | https://openalex.org/C104317684 |
| concepts[11].level | 2 |
| concepts[11].score | 0.24163460731506348 |
| concepts[11].wikidata | https://www.wikidata.org/wiki/Q7187 |
| concepts[11].display_name | Gene |
| keywords[0].id | https://openalex.org/keywords/genome-editing |
| keywords[0].score | 0.8775659799575806 |
| keywords[0].display_name | Genome editing |
| keywords[1].id | https://openalex.org/keywords/progenitor-cell |
| keywords[1].score | 0.667119026184082 |
| keywords[1].display_name | Progenitor cell |
| keywords[2].id | https://openalex.org/keywords/stem-cell |
| keywords[2].score | 0.5920475721359253 |
| keywords[2].display_name | Stem cell |
| keywords[3].id | https://openalex.org/keywords/computational-biology |
| keywords[3].score | 0.5532165765762329 |
| keywords[3].display_name | Computational biology |
| keywords[4].id | https://openalex.org/keywords/biology |
| keywords[4].score | 0.5474513173103333 |
| keywords[4].display_name | Biology |
| keywords[5].id | https://openalex.org/keywords/haematopoiesis |
| keywords[5].score | 0.5007486343383789 |
| keywords[5].display_name | Haematopoiesis |
| keywords[6].id | https://openalex.org/keywords/crispr |
| keywords[6].score | 0.4867730140686035 |
| keywords[6].display_name | CRISPR |
| keywords[7].id | https://openalex.org/keywords/cord-blood |
| keywords[7].score | 0.4371558129787445 |
| keywords[7].display_name | Cord blood |
| keywords[8].id | https://openalex.org/keywords/primary-cell |
| keywords[8].score | 0.41157066822052 |
| keywords[8].display_name | Primary cell |
| keywords[9].id | https://openalex.org/keywords/genetics |
| keywords[9].score | 0.3713715076446533 |
| keywords[9].display_name | Genetics |
| keywords[10].id | https://openalex.org/keywords/computer-science |
| keywords[10].score | 0.3530680537223816 |
| keywords[10].display_name | Computer science |
| keywords[11].id | https://openalex.org/keywords/gene |
| keywords[11].score | 0.24163460731506348 |
| keywords[11].display_name | Gene |
| language | en |
| locations[0].id | doi:10.7554/elife.91288.1.sa2 |
| locations[0].is_oa | True |
| locations[0].source | |
| locations[0].license | cc-by |
| locations[0].pdf_url | |
| locations[0].version | publishedVersion |
| locations[0].raw_type | peer-review |
| locations[0].license_id | https://openalex.org/licenses/cc-by |
| locations[0].is_accepted | True |
| locations[0].is_published | True |
| locations[0].raw_source_name | |
| locations[0].landing_page_url | https://doi.org/10.7554/elife.91288.1.sa2 |
| indexed_in | crossref |
| authorships[0].author.id | https://openalex.org/A5081633974 |
| authorships[0].author.orcid | https://orcid.org/0000-0002-0548-8477 |
| authorships[0].author.display_name | Jian Xu |
| authorships[0].author_position | first |
| authorships[0].raw_author_name | Jian Xu |
| authorships[0].is_corresponding | True |
| has_content.pdf | False |
| has_content.grobid_xml | False |
| is_paratext | False |
| open_access.is_oa | True |
| open_access.oa_url | https://doi.org/10.7554/elife.91288.1.sa2 |
| open_access.oa_status | gold |
| open_access.any_repository_has_fulltext | False |
| created_date | 2025-10-10T00:00:00 |
| display_name | eLife Assessment: Near-perfect precise on-target editing of human hematopoietic stem and progenitor cells |
| has_fulltext | False |
| is_retracted | False |
| updated_date | 2025-11-06T03:46:38.306776 |
| primary_topic.id | https://openalex.org/T10878 |
| primary_topic.field.id | https://openalex.org/fields/13 |
| primary_topic.field.display_name | Biochemistry, Genetics and Molecular Biology |
| primary_topic.score | 1.0 |
| primary_topic.domain.id | https://openalex.org/domains/1 |
| primary_topic.domain.display_name | Life Sciences |
| primary_topic.subfield.id | https://openalex.org/subfields/1312 |
| primary_topic.subfield.display_name | Molecular Biology |
| primary_topic.display_name | CRISPR and Genetic Engineering |
| related_works | https://openalex.org/W4366822718, https://openalex.org/W3030388918, https://openalex.org/W4200005163, https://openalex.org/W2807464733, https://openalex.org/W4220766053, https://openalex.org/W2603161682, https://openalex.org/W2960163646, https://openalex.org/W4213248511, https://openalex.org/W3165203721, https://openalex.org/W2991257722 |
| cited_by_count | 0 |
| locations_count | 1 |
| best_oa_location.id | doi:10.7554/elife.91288.1.sa2 |
| best_oa_location.is_oa | True |
| best_oa_location.source | |
| best_oa_location.license | cc-by |
| best_oa_location.pdf_url | |
| best_oa_location.version | publishedVersion |
| best_oa_location.raw_type | peer-review |
| best_oa_location.license_id | https://openalex.org/licenses/cc-by |
| best_oa_location.is_accepted | True |
| best_oa_location.is_published | True |
| best_oa_location.raw_source_name | |
| best_oa_location.landing_page_url | https://doi.org/10.7554/elife.91288.1.sa2 |
| primary_location.id | doi:10.7554/elife.91288.1.sa2 |
| primary_location.is_oa | True |
| primary_location.source | |
| primary_location.license | cc-by |
| primary_location.pdf_url | |
| primary_location.version | publishedVersion |
| primary_location.raw_type | peer-review |
| primary_location.license_id | https://openalex.org/licenses/cc-by |
| primary_location.is_accepted | True |
| primary_location.is_published | True |
| primary_location.raw_source_name | |
| primary_location.landing_page_url | https://doi.org/10.7554/elife.91288.1.sa2 |
| publication_date | 2023-10-05 |
| publication_year | 2023 |
| referenced_works_count | 0 |
| abstract_inverted_index.As | 126 |
| abstract_inverted_index.an | 35 |
| abstract_inverted_index.as | 19, 21 |
| abstract_inverted_index.at | 159 |
| abstract_inverted_index.be | 143, 164 |
| abstract_inverted_index.by | 145 |
| abstract_inverted_index.in | 3, 92, 95, 122, 147, 167, 175 |
| abstract_inverted_index.of | 37, 80, 88, 128, 150 |
| abstract_inverted_index.on | 54 |
| abstract_inverted_index.or | 118 |
| abstract_inverted_index.to | 72, 97 |
| abstract_inverted_index.we | 45, 133 |
| abstract_inverted_index.PAM | 101 |
| abstract_inverted_index.RNA | 39 |
| abstract_inverted_index.The | 67 |
| abstract_inverted_index.and | 7, 43, 62, 85, 111, 140, 152, 179 |
| abstract_inverted_index.can | 142, 162 |
| abstract_inverted_index.did | 112 |
| abstract_inverted_index.for | 16 |
| abstract_inverted_index.new | 173 |
| abstract_inverted_index.not | 113 |
| abstract_inverted_index.now | 47, 163 |
| abstract_inverted_index.the | 28, 78, 81, 86, 93, 100, 108, 116, 137 |
| abstract_inverted_index.was | 105 |
| abstract_inverted_index.>90% | 53 |
| abstract_inverted_index.This | 170 |
| abstract_inverted_index.With | 155 |
| abstract_inverted_index.also | 134 |
| abstract_inverted_index.both | 13, 176 |
| abstract_inverted_index.cell | 124 |
| abstract_inverted_index.cord | 56 |
| abstract_inverted_index.even | 26, 106 |
| abstract_inverted_index.gene | 1 |
| abstract_inverted_index.have | 46 |
| abstract_inverted_index.high | 75 |
| abstract_inverted_index.main | 68 |
| abstract_inverted_index.many | 129 |
| abstract_inverted_index.mean | 49 |
| abstract_inverted_index.open | 172 |
| abstract_inverted_index.stem | 6 |
| abstract_inverted_index.such | 74 |
| abstract_inverted_index.that | 136 |
| abstract_inverted_index.well | 20 |
| abstract_inverted_index.were | 77 |
| abstract_inverted_index.will | 171 |
| abstract_inverted_index.with | 27, 59 |
| abstract_inverted_index.HSCPs | 58 |
| abstract_inverted_index.blood | 57 |
| abstract_inverted_index.cells | 9 |
| abstract_inverted_index.donor | 41, 94 |
| abstract_inverted_index.guide | 38 |
| abstract_inverted_index.human | 168 |
| abstract_inverted_index.these | 156 |
| abstract_inverted_index.tuned | 144 |
| abstract_inverted_index.would | 11 |
| abstract_inverted_index.DNA-PK | 82 |
| abstract_inverted_index.HSPCs. | 169 |
| abstract_inverted_index.across | 107 |
| abstract_inverted_index.adding | 146 |
| abstract_inverted_index.affect | 119 |
| abstract_inverted_index.donor. | 154 |
| abstract_inverted_index.mutant | 151 |
| abstract_inverted_index.needed | 71 |
| abstract_inverted_index.remain | 32 |
| abstract_inverted_index.silent | 90 |
| abstract_inverted_index.(HSPCs) | 10 |
| abstract_inverted_index.Precise | 24 |
| abstract_inverted_index.Through | 34 |
| abstract_inverted_index.achieve | 73 |
| abstract_inverted_index.assays. | 125 |
| abstract_inverted_index.avenues | 174 |
| abstract_inverted_index.defined | 148 |
| abstract_inverted_index.design, | 42 |
| abstract_inverted_index.disease | 22, 180 |
| abstract_inverted_index.distort | 115 |
| abstract_inverted_index.editing | 2, 51, 104, 139, 158 |
| abstract_inverted_index.lineage | 120 |
| abstract_inverted_index.minimal | 60 |
| abstract_inverted_index.outputs | 121 |
| abstract_inverted_index.overall | 138 |
| abstract_inverted_index.precise | 50 |
| abstract_inverted_index.primary | 4, 55 |
| abstract_inverted_index.system, | 30 |
| abstract_inverted_index.without | 63 |
| abstract_inverted_index.AZD7648, | 84 |
| abstract_inverted_index.addition | 79, 96 |
| abstract_inverted_index.curative | 14 |
| abstract_inverted_index.directly | 166 |
| abstract_inverted_index.diseases | 130 |
| abstract_inverted_index.editing. | 66 |
| abstract_inverted_index.however, | 31 |
| abstract_inverted_index.limited. | 33 |
| abstract_inverted_index.mixtures | 149 |
| abstract_inverted_index.observed | 64 |
| abstract_inverted_index.obtained | 48 |
| abstract_inverted_index.protocol | 69 |
| abstract_inverted_index.requires | 131 |
| abstract_inverted_index.toxicity | 61 |
| abstract_inverted_index.zygosity | 141 |
| abstract_inverted_index.Precision | 0 |
| abstract_inverted_index.delivery, | 40 |
| abstract_inverted_index.disorders | 18 |
| abstract_inverted_index.hierarchy | 117 |
| abstract_inverted_index.inclusion | 87 |
| abstract_inverted_index.inhibitor | 83 |
| abstract_inverted_index.modelling | 127 |
| abstract_inverted_index.monogenic | 17 |
| abstract_inverted_index.mutations | 91, 98 |
| abstract_inverted_index.sequence. | 102 |
| abstract_inverted_index.wild-type | 153 |
| abstract_inverted_index.CRISPR/Cas | 29 |
| abstract_inverted_index.additives, | 44 |
| abstract_inverted_index.disrupting | 99 |
| abstract_inverted_index.efficiency | 161 |
| abstract_inverted_index.facilitate | 12 |
| abstract_inverted_index.hierarchy, | 110 |
| abstract_inverted_index.modelling. | 23, 181 |
| abstract_inverted_index.off-target | 65 |
| abstract_inverted_index.progenitor | 8, 109 |
| abstract_inverted_index.strategies | 178 |
| abstract_inverted_index.treatments | 15 |
| abstract_inverted_index.Critically, | 103 |
| abstract_inverted_index.therapeutic | 177 |
| abstract_inverted_index.accomplished | 165 |
| abstract_inverted_index.demonstrated | 135 |
| abstract_inverted_index.efficiencies | 25, 52, 76 |
| abstract_inverted_index.near-perfect | 160 |
| abstract_inverted_index.optimization | 36 |
| abstract_inverted_index.hematopoietic | 5 |
| abstract_inverted_index.modifications | 70 |
| abstract_inverted_index.substantially | 114 |
| abstract_inverted_index.colony-forming | 123 |
| abstract_inverted_index.optimizations, | 157 |
| abstract_inverted_index.heterozygosity, | 132 |
| abstract_inverted_index.spacer-breaking | 89 |
| cited_by_percentile_year | |
| corresponding_author_ids | https://openalex.org/A5081633974 |
| countries_distinct_count | 0 |
| institutions_distinct_count | 1 |
| sustainable_development_goals[0].id | https://metadata.un.org/sdg/3 |
| sustainable_development_goals[0].score | 0.7099999785423279 |
| sustainable_development_goals[0].display_name | Good health and well-being |
| citation_normalized_percentile |