Genetic dissection of microglia cannibalism reveals an IL10 signaling axis controls microglia lifespan Article Swipe
 YOU?
·
· 2025
· Open Access
·
· DOI: https://doi.org/10.1101/2025.11.21.689277
YOU?
·
· 2025
· Open Access
·
· DOI: https://doi.org/10.1101/2025.11.21.689277
The development of complex organs, like the brain, demands a robust system for tissue remodeling and cellular debris clearance. In the brain, this function is performed by microglia, which must clear diverse debris substrates, including that caused by cell death. Although the subsequent fate of these phagocytic microglia is a critical regulatory point that impacts whether the brain resolves a debris environment, the genetic mechanisms that control microglia fate after debris clearance remain mostly unknown. To address this, we conducted a large-scale CRISPR screen in zebrafish using a custom-built robotic confocal microscope. We selected candidate genes from a single-cell RNA sequencing dataset of embryonic mouse microglia. This screen identified several modulators of microglial lifespan and cannibalism that are enriched in mouse and zebrafish microglia, including interleukin-10 receptor beta ( il10rb ), a receptor subunit for the cytokine IL10. Perturbation of il10 , il10rb, and downstream signaling molecules JAK/STAT in zebrafish reduced microglial death. Expression analysis in mouse and zebrafish confirmed that microglia express both il10 and il10rb . Given the established role of IL10 in lysosomal remodeling, we hypothesized that it regulates microglial survival through lysosomal acidification. While il10rb perturbation did not alter lysosome number or size, it caused a significant reduction in LysoTracker-positive lysosomes, indicating decreased lysosomal acidification. Inhibiting v-ATPase also reduced microglial death, reinforcing the link between lysosomal pH and cell fate. Our findings reveal a cytokine-regulated mechanism where lysosomal dynamics determine the survival of phagocytic microglia. We propose that a necroptosis-cannibalism process functions as a quality control mechanism for microglial turnover, which is critical for refining neuroimmune cell function in the brain.
Related Topics
- Type
- article
- Landing Page
- https://doi.org/10.1101/2025.11.21.689277
- https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdf
- OA Status
- green
- References
- 66
- OpenAlex ID
- https://openalex.org/W7106286312
Raw OpenAlex JSON
- OpenAlex ID
-
https://openalex.org/W7106286312Canonical identifier for this work in OpenAlex
- DOI
-
https://doi.org/10.1101/2025.11.21.689277Digital Object Identifier
- Title
-
Genetic dissection of microglia cannibalism reveals an IL10 signaling axis controls microglia lifespanWork title
- Type
-
articleOpenAlex work type
- Publication year
-
2025Year of publication
- Publication date
-
2025-11-21Full publication date if available
- Authors
-
Hannah Gordon, Dailin Gan, Antonio Dolojan, John Dennen, Camden A. Hoover, Zachary M. Koh, Khang Chau, Ricky Avalos Arceo, Cara Cavanaugh, Jun Li, Cody J. Smith, Camden A. HooverList of authors in order
- Landing page
-
https://doi.org/10.1101/2025.11.21.689277Publisher landing page
- PDF URL
-
https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdfDirect link to full text PDF
- Open access
-
YesWhether a free full text is available
- OA status
-
greenOpen access status per OpenAlex
- OA URL
-
https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdfDirect OA link when available
- Concepts
-
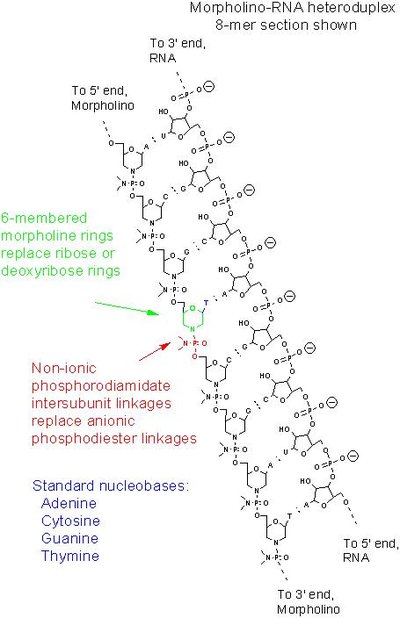
Microglia, Zebrafish, Biology, Cell biology, Lysosome, Receptor, Signal transduction, Embryonic stem cell, Model organism, Neuroscience, Cytokine, Fate mapping, Morpholino, Cell, Interleukin 10, Cell signaling, Central nervous system, Protein subunit, Gene, HEK 293 cells, Gene knockdown, Immune system, Immunology, Tumor necrosis factor alpha, Downregulation and upregulation, Growth cone, Cell type, Mechanism (biology), Notch signaling pathway, Cell fate determination, Regulation of gene expression, Genetic model, Nervous systemTop concepts (fields/topics) attached by OpenAlex
- Cited by
-
0Total citation count in OpenAlex
- References (count)
-
66Number of works referenced by this work
Full payload
| id | https://openalex.org/W7106286312 |
|---|---|
| doi | https://doi.org/10.1101/2025.11.21.689277 |
| ids.doi | https://doi.org/10.1101/2025.11.21.689277 |
| ids.openalex | https://openalex.org/W7106286312 |
| fwci | 0.0 |
| type | article |
| title | Genetic dissection of microglia cannibalism reveals an IL10 signaling axis controls microglia lifespan |
| biblio.issue | |
| biblio.volume | |
| biblio.last_page | |
| biblio.first_page | |
| topics[0].id | https://openalex.org/T11266 |
| topics[0].field.id | https://openalex.org/fields/28 |
| topics[0].field.display_name | Neuroscience |
| topics[0].score | 0.9771150350570679 |
| topics[0].domain.id | https://openalex.org/domains/1 |
| topics[0].domain.display_name | Life Sciences |
| topics[0].subfield.id | https://openalex.org/subfields/2808 |
| topics[0].subfield.display_name | Neurology |
| topics[0].display_name | Neuroinflammation and Neurodegeneration Mechanisms |
| topics[1].id | https://openalex.org/T11685 |
| topics[1].field.id | https://openalex.org/fields/13 |
| topics[1].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[1].score | 0.0107630779966712 |
| topics[1].domain.id | https://openalex.org/domains/1 |
| topics[1].domain.display_name | Life Sciences |
| topics[1].subfield.id | https://openalex.org/subfields/1307 |
| topics[1].subfield.display_name | Cell Biology |
| topics[1].display_name | Zebrafish Biomedical Research Applications |
| topics[2].id | https://openalex.org/T12672 |
| topics[2].field.id | https://openalex.org/fields/24 |
| topics[2].field.display_name | Immunology and Microbiology |
| topics[2].score | 0.0027170530520379543 |
| topics[2].domain.id | https://openalex.org/domains/1 |
| topics[2].domain.display_name | Life Sciences |
| topics[2].subfield.id | https://openalex.org/subfields/2403 |
| topics[2].subfield.display_name | Immunology |
| topics[2].display_name | Phagocytosis and Immune Regulation |
| is_xpac | False |
| apc_list | |
| apc_paid | |
| concepts[0].id | https://openalex.org/C2779830541 |
| concepts[0].level | 3 |
| concepts[0].score | 0.9032859206199646 |
| concepts[0].wikidata | https://www.wikidata.org/wiki/Q1622829 |
| concepts[0].display_name | Microglia |
| concepts[1].id | https://openalex.org/C2776878037 |
| concepts[1].level | 3 |
| concepts[1].score | 0.8407309651374817 |
| concepts[1].wikidata | https://www.wikidata.org/wiki/Q169444 |
| concepts[1].display_name | Zebrafish |
| concepts[2].id | https://openalex.org/C86803240 |
| concepts[2].level | 0 |
| concepts[2].score | 0.7898026704788208 |
| concepts[2].wikidata | https://www.wikidata.org/wiki/Q420 |
| concepts[2].display_name | Biology |
| concepts[3].id | https://openalex.org/C95444343 |
| concepts[3].level | 1 |
| concepts[3].score | 0.7011377215385437 |
| concepts[3].wikidata | https://www.wikidata.org/wiki/Q7141 |
| concepts[3].display_name | Cell biology |
| concepts[4].id | https://openalex.org/C2780216420 |
| concepts[4].level | 3 |
| concepts[4].score | 0.5009453892707825 |
| concepts[4].wikidata | https://www.wikidata.org/wiki/Q83330 |
| concepts[4].display_name | Lysosome |
| concepts[5].id | https://openalex.org/C170493617 |
| concepts[5].level | 2 |
| concepts[5].score | 0.4623804986476898 |
| concepts[5].wikidata | https://www.wikidata.org/wiki/Q208467 |
| concepts[5].display_name | Receptor |
| concepts[6].id | https://openalex.org/C62478195 |
| concepts[6].level | 2 |
| concepts[6].score | 0.4377354383468628 |
| concepts[6].wikidata | https://www.wikidata.org/wiki/Q828130 |
| concepts[6].display_name | Signal transduction |
| concepts[7].id | https://openalex.org/C145103041 |
| concepts[7].level | 3 |
| concepts[7].score | 0.43226194381713867 |
| concepts[7].wikidata | https://www.wikidata.org/wiki/Q1151519 |
| concepts[7].display_name | Embryonic stem cell |
| concepts[8].id | https://openalex.org/C19843653 |
| concepts[8].level | 3 |
| concepts[8].score | 0.42309215664863586 |
| concepts[8].wikidata | https://www.wikidata.org/wiki/Q213907 |
| concepts[8].display_name | Model organism |
| concepts[9].id | https://openalex.org/C169760540 |
| concepts[9].level | 1 |
| concepts[9].score | 0.3558383285999298 |
| concepts[9].wikidata | https://www.wikidata.org/wiki/Q207011 |
| concepts[9].display_name | Neuroscience |
| concepts[10].id | https://openalex.org/C2778690821 |
| concepts[10].level | 2 |
| concepts[10].score | 0.34787717461586 |
| concepts[10].wikidata | https://www.wikidata.org/wiki/Q212354 |
| concepts[10].display_name | Cytokine |
| concepts[11].id | https://openalex.org/C63101894 |
| concepts[11].level | 4 |
| concepts[11].score | 0.3464736342430115 |
| concepts[11].wikidata | https://www.wikidata.org/wiki/Q604095 |
| concepts[11].display_name | Fate mapping |
| concepts[12].id | https://openalex.org/C136571751 |
| concepts[12].level | 4 |
| concepts[12].score | 0.3424054980278015 |
| concepts[12].wikidata | https://www.wikidata.org/wiki/Q424607 |
| concepts[12].display_name | Morpholino |
| concepts[13].id | https://openalex.org/C1491633281 |
| concepts[13].level | 2 |
| concepts[13].score | 0.33870363235473633 |
| concepts[13].wikidata | https://www.wikidata.org/wiki/Q7868 |
| concepts[13].display_name | Cell |
| concepts[14].id | https://openalex.org/C187316574 |
| concepts[14].level | 3 |
| concepts[14].score | 0.3181261122226715 |
| concepts[14].wikidata | https://www.wikidata.org/wiki/Q423698 |
| concepts[14].display_name | Interleukin 10 |
| concepts[15].id | https://openalex.org/C150555746 |
| concepts[15].level | 3 |
| concepts[15].score | 0.31351667642593384 |
| concepts[15].wikidata | https://www.wikidata.org/wiki/Q210973 |
| concepts[15].display_name | Cell signaling |
| concepts[16].id | https://openalex.org/C529278444 |
| concepts[16].level | 2 |
| concepts[16].score | 0.29693347215652466 |
| concepts[16].wikidata | https://www.wikidata.org/wiki/Q47273 |
| concepts[16].display_name | Central nervous system |
| concepts[17].id | https://openalex.org/C104292427 |
| concepts[17].level | 3 |
| concepts[17].score | 0.29500725865364075 |
| concepts[17].wikidata | https://www.wikidata.org/wiki/Q899781 |
| concepts[17].display_name | Protein subunit |
| concepts[18].id | https://openalex.org/C104317684 |
| concepts[18].level | 2 |
| concepts[18].score | 0.29134610295295715 |
| concepts[18].wikidata | https://www.wikidata.org/wiki/Q7187 |
| concepts[18].display_name | Gene |
| concepts[19].id | https://openalex.org/C174510640 |
| concepts[19].level | 3 |
| concepts[19].score | 0.28179892897605896 |
| concepts[19].wikidata | https://www.wikidata.org/wiki/Q489618 |
| concepts[19].display_name | HEK 293 cells |
| concepts[20].id | https://openalex.org/C173396325 |
| concepts[20].level | 3 |
| concepts[20].score | 0.2791925370693207 |
| concepts[20].wikidata | https://www.wikidata.org/wiki/Q1501275 |
| concepts[20].display_name | Gene knockdown |
| concepts[21].id | https://openalex.org/C8891405 |
| concepts[21].level | 2 |
| concepts[21].score | 0.27704787254333496 |
| concepts[21].wikidata | https://www.wikidata.org/wiki/Q1059 |
| concepts[21].display_name | Immune system |
| concepts[22].id | https://openalex.org/C203014093 |
| concepts[22].level | 1 |
| concepts[22].score | 0.27521708607673645 |
| concepts[22].wikidata | https://www.wikidata.org/wiki/Q101929 |
| concepts[22].display_name | Immunology |
| concepts[23].id | https://openalex.org/C17991360 |
| concepts[23].level | 2 |
| concepts[23].score | 0.27470317482948303 |
| concepts[23].wikidata | https://www.wikidata.org/wiki/Q21173843 |
| concepts[23].display_name | Tumor necrosis factor alpha |
| concepts[24].id | https://openalex.org/C127561419 |
| concepts[24].level | 3 |
| concepts[24].score | 0.27263960242271423 |
| concepts[24].wikidata | https://www.wikidata.org/wiki/Q224527 |
| concepts[24].display_name | Downregulation and upregulation |
| concepts[25].id | https://openalex.org/C135903288 |
| concepts[25].level | 3 |
| concepts[25].score | 0.2720628082752228 |
| concepts[25].wikidata | https://www.wikidata.org/wiki/Q559423 |
| concepts[25].display_name | Growth cone |
| concepts[26].id | https://openalex.org/C189014844 |
| concepts[26].level | 3 |
| concepts[26].score | 0.2718566954135895 |
| concepts[26].wikidata | https://www.wikidata.org/wiki/Q189118 |
| concepts[26].display_name | Cell type |
| concepts[27].id | https://openalex.org/C89611455 |
| concepts[27].level | 2 |
| concepts[27].score | 0.26695534586906433 |
| concepts[27].wikidata | https://www.wikidata.org/wiki/Q6804646 |
| concepts[27].display_name | Mechanism (biology) |
| concepts[28].id | https://openalex.org/C161879069 |
| concepts[28].level | 3 |
| concepts[28].score | 0.26634347438812256 |
| concepts[28].wikidata | https://www.wikidata.org/wiki/Q904082 |
| concepts[28].display_name | Notch signaling pathway |
| concepts[29].id | https://openalex.org/C163952510 |
| concepts[29].level | 4 |
| concepts[29].score | 0.2653423249721527 |
| concepts[29].wikidata | https://www.wikidata.org/wiki/Q5058196 |
| concepts[29].display_name | Cell fate determination |
| concepts[30].id | https://openalex.org/C165864922 |
| concepts[30].level | 3 |
| concepts[30].score | 0.25655993819236755 |
| concepts[30].wikidata | https://www.wikidata.org/wiki/Q411391 |
| concepts[30].display_name | Regulation of gene expression |
| concepts[31].id | https://openalex.org/C2992519594 |
| concepts[31].level | 3 |
| concepts[31].score | 0.2559390664100647 |
| concepts[31].wikidata | https://www.wikidata.org/wiki/Q870433 |
| concepts[31].display_name | Genetic model |
| concepts[32].id | https://openalex.org/C545706735 |
| concepts[32].level | 2 |
| concepts[32].score | 0.2500099539756775 |
| concepts[32].wikidata | https://www.wikidata.org/wiki/Q9404 |
| concepts[32].display_name | Nervous system |
| keywords[0].id | https://openalex.org/keywords/microglia |
| keywords[0].score | 0.9032859206199646 |
| keywords[0].display_name | Microglia |
| keywords[1].id | https://openalex.org/keywords/zebrafish |
| keywords[1].score | 0.8407309651374817 |
| keywords[1].display_name | Zebrafish |
| keywords[2].id | https://openalex.org/keywords/lysosome |
| keywords[2].score | 0.5009453892707825 |
| keywords[2].display_name | Lysosome |
| keywords[3].id | https://openalex.org/keywords/receptor |
| keywords[3].score | 0.4623804986476898 |
| keywords[3].display_name | Receptor |
| keywords[4].id | https://openalex.org/keywords/signal-transduction |
| keywords[4].score | 0.4377354383468628 |
| keywords[4].display_name | Signal transduction |
| keywords[5].id | https://openalex.org/keywords/embryonic-stem-cell |
| keywords[5].score | 0.43226194381713867 |
| keywords[5].display_name | Embryonic stem cell |
| keywords[6].id | https://openalex.org/keywords/model-organism |
| keywords[6].score | 0.42309215664863586 |
| keywords[6].display_name | Model organism |
| keywords[7].id | https://openalex.org/keywords/cytokine |
| keywords[7].score | 0.34787717461586 |
| keywords[7].display_name | Cytokine |
| keywords[8].id | https://openalex.org/keywords/fate-mapping |
| keywords[8].score | 0.3464736342430115 |
| keywords[8].display_name | Fate mapping |
| keywords[9].id | https://openalex.org/keywords/morpholino |
| keywords[9].score | 0.3424054980278015 |
| keywords[9].display_name | Morpholino |
| language | |
| locations[0].id | doi:10.1101/2025.11.21.689277 |
| locations[0].is_oa | True |
| locations[0].source.id | https://openalex.org/S4306402567 |
| locations[0].source.issn | |
| locations[0].source.type | repository |
| locations[0].source.is_oa | False |
| locations[0].source.issn_l | |
| locations[0].source.is_core | False |
| locations[0].source.is_in_doaj | False |
| locations[0].source.display_name | bioRxiv (Cold Spring Harbor Laboratory) |
| locations[0].source.host_organization | https://openalex.org/I2750212522 |
| locations[0].source.host_organization_name | Cold Spring Harbor Laboratory |
| locations[0].source.host_organization_lineage | https://openalex.org/I2750212522 |
| locations[0].license | |
| locations[0].pdf_url | https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdf |
| locations[0].version | acceptedVersion |
| locations[0].raw_type | posted-content |
| locations[0].license_id | |
| locations[0].is_accepted | True |
| locations[0].is_published | False |
| locations[0].raw_source_name | |
| locations[0].landing_page_url | https://doi.org/10.1101/2025.11.21.689277 |
| indexed_in | crossref |
| authorships[0].author.id | https://openalex.org/A2129145359 |
| authorships[0].author.orcid | https://orcid.org/0000-0001-7510-0071 |
| authorships[0].author.display_name | Hannah Gordon |
| authorships[0].countries | US |
| authorships[0].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[0].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[0].institutions[0].id | https://openalex.org/I107639228 |
| authorships[0].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[0].institutions[0].type | education |
| authorships[0].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[0].institutions[0].country_code | US |
| authorships[0].institutions[0].display_name | University of Notre Dame |
| authorships[0].author_position | first |
| authorships[0].raw_author_name | Hannah Gordon |
| authorships[0].is_corresponding | False |
| authorships[0].raw_affiliation_strings | University of Notre Dame |
| authorships[1].author.id | https://openalex.org/A3125025023 |
| authorships[1].author.orcid | https://orcid.org/0000-0003-0535-1259 |
| authorships[1].author.display_name | Dailin Gan |
| authorships[1].countries | US |
| authorships[1].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[1].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[1].institutions[0].id | https://openalex.org/I107639228 |
| authorships[1].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[1].institutions[0].type | education |
| authorships[1].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[1].institutions[0].country_code | US |
| authorships[1].institutions[0].display_name | University of Notre Dame |
| authorships[1].author_position | middle |
| authorships[1].raw_author_name | Dailin Gan |
| authorships[1].is_corresponding | False |
| authorships[1].raw_affiliation_strings | University of Notre Dame |
| authorships[2].author.id | https://openalex.org/A5116551099 |
| authorships[2].author.orcid | |
| authorships[2].author.display_name | Antonio Dolojan |
| authorships[2].countries | US |
| authorships[2].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[2].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[2].institutions[0].id | https://openalex.org/I107639228 |
| authorships[2].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[2].institutions[0].type | education |
| authorships[2].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[2].institutions[0].country_code | US |
| authorships[2].institutions[0].display_name | University of Notre Dame |
| authorships[2].author_position | middle |
| authorships[2].raw_author_name | Antonio Dolojan |
| authorships[2].is_corresponding | False |
| authorships[2].raw_affiliation_strings | University of Notre Dame |
| authorships[3].author.id | https://openalex.org/A5119141195 |
| authorships[3].author.orcid | |
| authorships[3].author.display_name | John Dennen |
| authorships[3].countries | US |
| authorships[3].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[3].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[3].institutions[0].id | https://openalex.org/I107639228 |
| authorships[3].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[3].institutions[0].type | education |
| authorships[3].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[3].institutions[0].country_code | US |
| authorships[3].institutions[0].display_name | University of Notre Dame |
| authorships[3].author_position | middle |
| authorships[3].raw_author_name | John Dennen |
| authorships[3].is_corresponding | False |
| authorships[3].raw_affiliation_strings | University of Notre Dame |
| authorships[4].author.id | https://openalex.org/A2979418281 |
| authorships[4].author.orcid | |
| authorships[4].author.display_name | Camden A. Hoover |
| authorships[4].countries | US |
| authorships[4].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[4].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[4].institutions[0].id | https://openalex.org/I107639228 |
| authorships[4].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[4].institutions[0].type | education |
| authorships[4].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[4].institutions[0].country_code | US |
| authorships[4].institutions[0].display_name | University of Notre Dame |
| authorships[4].author_position | middle |
| authorships[4].raw_author_name | Camden A. Hoover |
| authorships[4].is_corresponding | False |
| authorships[4].raw_affiliation_strings | University of Notre Dame |
| authorships[5].author.id | https://openalex.org/A5116551098 |
| authorships[5].author.orcid | |
| authorships[5].author.display_name | Zachary M. Koh |
| authorships[5].countries | US |
| authorships[5].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[5].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[5].institutions[0].id | https://openalex.org/I107639228 |
| authorships[5].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[5].institutions[0].type | education |
| authorships[5].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[5].institutions[0].country_code | US |
| authorships[5].institutions[0].display_name | University of Notre Dame |
| authorships[5].author_position | middle |
| authorships[5].raw_author_name | Zachary M. Koh |
| authorships[5].is_corresponding | False |
| authorships[5].raw_affiliation_strings | University of Notre Dame |
| authorships[6].author.id | https://openalex.org/A2790787785 |
| authorships[6].author.orcid | https://orcid.org/0000-0002-9112-2470 |
| authorships[6].author.display_name | Khang Chau |
| authorships[6].countries | US |
| authorships[6].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[6].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[6].institutions[0].id | https://openalex.org/I107639228 |
| authorships[6].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[6].institutions[0].type | education |
| authorships[6].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[6].institutions[0].country_code | US |
| authorships[6].institutions[0].display_name | University of Notre Dame |
| authorships[6].author_position | middle |
| authorships[6].raw_author_name | Khang Chau |
| authorships[6].is_corresponding | False |
| authorships[6].raw_affiliation_strings | University of Notre Dame |
| authorships[7].author.id | https://openalex.org/A5076471094 |
| authorships[7].author.orcid | |
| authorships[7].author.display_name | Ricky Avalos Arceo |
| authorships[7].countries | US |
| authorships[7].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[7].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[7].institutions[0].id | https://openalex.org/I107639228 |
| authorships[7].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[7].institutions[0].type | education |
| authorships[7].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[7].institutions[0].country_code | US |
| authorships[7].institutions[0].display_name | University of Notre Dame |
| authorships[7].author_position | middle |
| authorships[7].raw_author_name | Ricky Avalos Arceo |
| authorships[7].is_corresponding | False |
| authorships[7].raw_affiliation_strings | University of Notre Dame |
| authorships[8].author.id | https://openalex.org/A3188820470 |
| authorships[8].author.orcid | |
| authorships[8].author.display_name | Cara Cavanaugh |
| authorships[8].countries | US |
| authorships[8].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[8].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[8].institutions[0].id | https://openalex.org/I107639228 |
| authorships[8].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[8].institutions[0].type | education |
| authorships[8].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[8].institutions[0].country_code | US |
| authorships[8].institutions[0].display_name | University of Notre Dame |
| authorships[8].author_position | middle |
| authorships[8].raw_author_name | Cara Cavanaugh |
| authorships[8].is_corresponding | False |
| authorships[8].raw_affiliation_strings | University of Notre Dame |
| authorships[9].author.id | https://openalex.org/A2012836890 |
| authorships[9].author.orcid | https://orcid.org/0000-0001-5027-1624 |
| authorships[9].author.display_name | Jun Li |
| authorships[9].countries | US |
| authorships[9].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[9].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[9].institutions[0].id | https://openalex.org/I107639228 |
| authorships[9].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[9].institutions[0].type | education |
| authorships[9].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[9].institutions[0].country_code | US |
| authorships[9].institutions[0].display_name | University of Notre Dame |
| authorships[9].author_position | middle |
| authorships[9].raw_author_name | Jun Li |
| authorships[9].is_corresponding | False |
| authorships[9].raw_affiliation_strings | University of Notre Dame |
| authorships[10].author.id | https://openalex.org/A2639751708 |
| authorships[10].author.orcid | https://orcid.org/0000-0002-9831-1514 |
| authorships[10].author.display_name | Cody J. Smith |
| authorships[10].countries | US |
| authorships[10].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[10].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[10].institutions[0].id | https://openalex.org/I107639228 |
| authorships[10].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[10].institutions[0].type | education |
| authorships[10].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[10].institutions[0].country_code | US |
| authorships[10].institutions[0].display_name | University of Notre Dame |
| authorships[10].author_position | middle |
| authorships[10].raw_author_name | Cody J. Smith |
| authorships[10].is_corresponding | True |
| authorships[10].raw_affiliation_strings | University of Notre Dame |
| authorships[11].author.id | https://openalex.org/A2979418281 |
| authorships[11].author.orcid | |
| authorships[11].author.display_name | Camden A. Hoover |
| authorships[11].countries | US |
| authorships[11].affiliations[0].institution_ids | https://openalex.org/I107639228 |
| authorships[11].affiliations[0].raw_affiliation_string | University of Notre Dame |
| authorships[11].institutions[0].id | https://openalex.org/I107639228 |
| authorships[11].institutions[0].ror | https://ror.org/https://ror.org/00mkhxb43 |
| authorships[11].institutions[0].type | education |
| authorships[11].institutions[0].lineage | https://openalex.org/I107639228 |
| authorships[11].institutions[0].country_code | US |
| authorships[11].institutions[0].display_name | University of Notre Dame |
| authorships[11].author_position | last |
| authorships[11].raw_author_name | Camden A Hoover |
| authorships[11].is_corresponding | False |
| authorships[11].raw_affiliation_strings | University of Notre Dame |
| has_content.pdf | True |
| has_content.grobid_xml | False |
| is_paratext | False |
| open_access.is_oa | True |
| open_access.oa_url | https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdf |
| open_access.oa_status | green |
| open_access.any_repository_has_fulltext | False |
| created_date | 2025-11-23T00:00:00 |
| display_name | Genetic dissection of microglia cannibalism reveals an IL10 signaling axis controls microglia lifespan |
| has_fulltext | False |
| is_retracted | False |
| updated_date | 2025-11-25T14:43:58.451035 |
| primary_topic.id | https://openalex.org/T11266 |
| primary_topic.field.id | https://openalex.org/fields/28 |
| primary_topic.field.display_name | Neuroscience |
| primary_topic.score | 0.9771150350570679 |
| primary_topic.domain.id | https://openalex.org/domains/1 |
| primary_topic.domain.display_name | Life Sciences |
| primary_topic.subfield.id | https://openalex.org/subfields/2808 |
| primary_topic.subfield.display_name | Neurology |
| primary_topic.display_name | Neuroinflammation and Neurodegeneration Mechanisms |
| cited_by_count | 0 |
| locations_count | 1 |
| best_oa_location.id | doi:10.1101/2025.11.21.689277 |
| best_oa_location.is_oa | True |
| best_oa_location.source.id | https://openalex.org/S4306402567 |
| best_oa_location.source.issn | |
| best_oa_location.source.type | repository |
| best_oa_location.source.is_oa | False |
| best_oa_location.source.issn_l | |
| best_oa_location.source.is_core | False |
| best_oa_location.source.is_in_doaj | False |
| best_oa_location.source.display_name | bioRxiv (Cold Spring Harbor Laboratory) |
| best_oa_location.source.host_organization | https://openalex.org/I2750212522 |
| best_oa_location.source.host_organization_name | Cold Spring Harbor Laboratory |
| best_oa_location.source.host_organization_lineage | https://openalex.org/I2750212522 |
| best_oa_location.license | |
| best_oa_location.pdf_url | https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdf |
| best_oa_location.version | acceptedVersion |
| best_oa_location.raw_type | posted-content |
| best_oa_location.license_id | |
| best_oa_location.is_accepted | True |
| best_oa_location.is_published | False |
| best_oa_location.raw_source_name | |
| best_oa_location.landing_page_url | https://doi.org/10.1101/2025.11.21.689277 |
| primary_location.id | doi:10.1101/2025.11.21.689277 |
| primary_location.is_oa | True |
| primary_location.source.id | https://openalex.org/S4306402567 |
| primary_location.source.issn | |
| primary_location.source.type | repository |
| primary_location.source.is_oa | False |
| primary_location.source.issn_l | |
| primary_location.source.is_core | False |
| primary_location.source.is_in_doaj | False |
| primary_location.source.display_name | bioRxiv (Cold Spring Harbor Laboratory) |
| primary_location.source.host_organization | https://openalex.org/I2750212522 |
| primary_location.source.host_organization_name | Cold Spring Harbor Laboratory |
| primary_location.source.host_organization_lineage | https://openalex.org/I2750212522 |
| primary_location.license | |
| primary_location.pdf_url | https://www.biorxiv.org/content/biorxiv/early/2025/11/21/2025.11.21.689277.full.pdf |
| primary_location.version | acceptedVersion |
| primary_location.raw_type | posted-content |
| primary_location.license_id | |
| primary_location.is_accepted | True |
| primary_location.is_published | False |
| primary_location.raw_source_name | |
| primary_location.landing_page_url | https://doi.org/10.1101/2025.11.21.689277 |
| publication_date | 2025-11-21 |
| publication_year | 2025 |
| referenced_works | https://openalex.org/W4308150510, https://openalex.org/W2492998259, https://openalex.org/W3013366780, https://openalex.org/W4225795175, https://openalex.org/W2029190354, https://openalex.org/W2151844149, https://openalex.org/W2088350077, https://openalex.org/W4200290917, https://openalex.org/W4309492951, https://openalex.org/W4391818018, https://openalex.org/W3013196639, https://openalex.org/W3015080429, https://openalex.org/W2053603253, https://openalex.org/W2888210447, https://openalex.org/W2402129233, https://openalex.org/W3087073735, https://openalex.org/W4281737563, https://openalex.org/W3010985052, https://openalex.org/W4414103841, https://openalex.org/W4408061745, https://openalex.org/W2052228332, https://openalex.org/W3090732972, https://openalex.org/W2044448807, https://openalex.org/W4393016060, https://openalex.org/W4390507681, https://openalex.org/W2040719456, https://openalex.org/W4225585633, https://openalex.org/W2057258619, https://openalex.org/W4403997658, https://openalex.org/W2113934014, https://openalex.org/W4220775671, https://openalex.org/W2007106585, https://openalex.org/W4403920202, https://openalex.org/W2901838561, https://openalex.org/W1964982794, https://openalex.org/W3092058471, https://openalex.org/W4220721798, https://openalex.org/W2164232024, https://openalex.org/W4399143068, https://openalex.org/W2025144487, https://openalex.org/W2082124619, https://openalex.org/W2099558832, https://openalex.org/W2888443298, https://openalex.org/W3175857880, https://openalex.org/W2766243612, https://openalex.org/W4283027720, https://openalex.org/W4280512820, https://openalex.org/W4220737780, https://openalex.org/W2148898943, https://openalex.org/W4415407394, https://openalex.org/W3216177033, https://openalex.org/W2025700341, https://openalex.org/W2972796665, https://openalex.org/W4307340212, https://openalex.org/W2110796758, https://openalex.org/W2000678720, https://openalex.org/W2888292224, https://openalex.org/W2070217465, https://openalex.org/W2070740895, https://openalex.org/W4206577698, https://openalex.org/W2075525665, https://openalex.org/W2125349610, https://openalex.org/W4412804184, https://openalex.org/W2911250221, https://openalex.org/W3214120156, https://openalex.org/W2912245056 |
| referenced_works_count | 66 |
| abstract_inverted_index.( | 129 |
| abstract_inverted_index., | 142 |
| abstract_inverted_index.. | 168 |
| abstract_inverted_index.a | 10, 50, 60, 81, 88, 98, 132, 200, 228, 243, 248 |
| abstract_inverted_index.), | 131 |
| abstract_inverted_index.In | 20 |
| abstract_inverted_index.To | 76 |
| abstract_inverted_index.We | 93, 240 |
| abstract_inverted_index.as | 247 |
| abstract_inverted_index.by | 27, 38 |
| abstract_inverted_index.in | 85, 120, 149, 156, 175, 203, 263 |
| abstract_inverted_index.is | 25, 49, 256 |
| abstract_inverted_index.it | 181, 198 |
| abstract_inverted_index.of | 3, 45, 103, 112, 140, 173, 237 |
| abstract_inverted_index.or | 196 |
| abstract_inverted_index.pH | 221 |
| abstract_inverted_index.we | 79, 178 |
| abstract_inverted_index.Our | 225 |
| abstract_inverted_index.RNA | 100 |
| abstract_inverted_index.The | 1 |
| abstract_inverted_index.and | 16, 115, 122, 144, 158, 166, 222 |
| abstract_inverted_index.are | 118 |
| abstract_inverted_index.did | 191 |
| abstract_inverted_index.for | 13, 135, 252, 258 |
| abstract_inverted_index.not | 192 |
| abstract_inverted_index.the | 7, 21, 42, 57, 63, 136, 170, 217, 235, 264 |
| abstract_inverted_index.IL10 | 174 |
| abstract_inverted_index.This | 107 |
| abstract_inverted_index.also | 212 |
| abstract_inverted_index.beta | 128 |
| abstract_inverted_index.both | 164 |
| abstract_inverted_index.cell | 39, 223, 261 |
| abstract_inverted_index.fate | 44, 69 |
| abstract_inverted_index.from | 97 |
| abstract_inverted_index.il10 | 141, 165 |
| abstract_inverted_index.like | 6 |
| abstract_inverted_index.link | 218 |
| abstract_inverted_index.must | 30 |
| abstract_inverted_index.role | 172 |
| abstract_inverted_index.that | 36, 54, 66, 117, 161, 180, 242 |
| abstract_inverted_index.this | 23 |
| abstract_inverted_index.Given | 169 |
| abstract_inverted_index.IL10. | 138 |
| abstract_inverted_index.While | 188 |
| abstract_inverted_index.after | 70 |
| abstract_inverted_index.alter | 193 |
| abstract_inverted_index.brain | 58 |
| abstract_inverted_index.clear | 31 |
| abstract_inverted_index.fate. | 224 |
| abstract_inverted_index.genes | 96 |
| abstract_inverted_index.mouse | 105, 121, 157 |
| abstract_inverted_index.point | 53 |
| abstract_inverted_index.size, | 197 |
| abstract_inverted_index.these | 46 |
| abstract_inverted_index.this, | 78 |
| abstract_inverted_index.using | 87 |
| abstract_inverted_index.where | 231 |
| abstract_inverted_index.which | 29, 255 |
| abstract_inverted_index.CRISPR | 83 |
| abstract_inverted_index.brain, | 8, 22 |
| abstract_inverted_index.brain. | 265 |
| abstract_inverted_index.caused | 37, 199 |
| abstract_inverted_index.death, | 215 |
| abstract_inverted_index.death. | 40, 153 |
| abstract_inverted_index.debris | 18, 33, 61, 71 |
| abstract_inverted_index.il10rb | 130, 167, 189 |
| abstract_inverted_index.mostly | 74 |
| abstract_inverted_index.number | 195 |
| abstract_inverted_index.remain | 73 |
| abstract_inverted_index.reveal | 227 |
| abstract_inverted_index.robust | 11 |
| abstract_inverted_index.screen | 84, 108 |
| abstract_inverted_index.system | 12 |
| abstract_inverted_index.tissue | 14 |
| abstract_inverted_index.address | 77 |
| abstract_inverted_index.between | 219 |
| abstract_inverted_index.complex | 4 |
| abstract_inverted_index.control | 67, 250 |
| abstract_inverted_index.dataset | 102 |
| abstract_inverted_index.demands | 9 |
| abstract_inverted_index.diverse | 32 |
| abstract_inverted_index.express | 163 |
| abstract_inverted_index.genetic | 64 |
| abstract_inverted_index.il10rb, | 143 |
| abstract_inverted_index.impacts | 55 |
| abstract_inverted_index.organs, | 5 |
| abstract_inverted_index.process | 245 |
| abstract_inverted_index.propose | 241 |
| abstract_inverted_index.quality | 249 |
| abstract_inverted_index.reduced | 151, 213 |
| abstract_inverted_index.robotic | 90 |
| abstract_inverted_index.several | 110 |
| abstract_inverted_index.subunit | 134 |
| abstract_inverted_index.through | 185 |
| abstract_inverted_index.whether | 56 |
| abstract_inverted_index.Abstract | 0 |
| abstract_inverted_index.Although | 41 |
| abstract_inverted_index.JAK/STAT | 148 |
| abstract_inverted_index.analysis | 155 |
| abstract_inverted_index.cellular | 17 |
| abstract_inverted_index.confocal | 91 |
| abstract_inverted_index.critical | 51, 257 |
| abstract_inverted_index.cytokine | 137 |
| abstract_inverted_index.dynamics | 233 |
| abstract_inverted_index.enriched | 119 |
| abstract_inverted_index.findings | 226 |
| abstract_inverted_index.function | 24, 262 |
| abstract_inverted_index.lifespan | 114 |
| abstract_inverted_index.lysosome | 194 |
| abstract_inverted_index.receptor | 127, 133 |
| abstract_inverted_index.refining | 259 |
| abstract_inverted_index.resolves | 59 |
| abstract_inverted_index.selected | 94 |
| abstract_inverted_index.survival | 184, 236 |
| abstract_inverted_index.unknown. | 75 |
| abstract_inverted_index.v-ATPase | 211 |
| abstract_inverted_index.candidate | 95 |
| abstract_inverted_index.clearance | 72 |
| abstract_inverted_index.conducted | 80 |
| abstract_inverted_index.confirmed | 160 |
| abstract_inverted_index.decreased | 207 |
| abstract_inverted_index.determine | 234 |
| abstract_inverted_index.embryonic | 104 |
| abstract_inverted_index.functions | 246 |
| abstract_inverted_index.including | 35, 125 |
| abstract_inverted_index.lysosomal | 176, 186, 208, 220, 232 |
| abstract_inverted_index.mechanism | 230, 251 |
| abstract_inverted_index.microglia | 48, 68, 162 |
| abstract_inverted_index.molecules | 147 |
| abstract_inverted_index.performed | 26 |
| abstract_inverted_index.reduction | 202 |
| abstract_inverted_index.regulates | 182 |
| abstract_inverted_index.signaling | 146 |
| abstract_inverted_index.turnover, | 254 |
| abstract_inverted_index.zebrafish | 86, 123, 150, 159 |
| abstract_inverted_index.Expression | 154 |
| abstract_inverted_index.Inhibiting | 210 |
| abstract_inverted_index.clearance. | 19 |
| abstract_inverted_index.downstream | 145 |
| abstract_inverted_index.identified | 109 |
| abstract_inverted_index.indicating | 206 |
| abstract_inverted_index.lysosomes, | 205 |
| abstract_inverted_index.mechanisms | 65 |
| abstract_inverted_index.microglia, | 28, 124 |
| abstract_inverted_index.microglia. | 106, 239 |
| abstract_inverted_index.microglial | 113, 152, 183, 214, 253 |
| abstract_inverted_index.modulators | 111 |
| abstract_inverted_index.phagocytic | 47, 238 |
| abstract_inverted_index.regulatory | 52 |
| abstract_inverted_index.remodeling | 15 |
| abstract_inverted_index.sequencing | 101 |
| abstract_inverted_index.subsequent | 43 |
| abstract_inverted_index.cannibalism | 116 |
| abstract_inverted_index.development | 2 |
| abstract_inverted_index.established | 171 |
| abstract_inverted_index.large-scale | 82 |
| abstract_inverted_index.microscope. | 92 |
| abstract_inverted_index.neuroimmune | 260 |
| abstract_inverted_index.reinforcing | 216 |
| abstract_inverted_index.remodeling, | 177 |
| abstract_inverted_index.significant | 201 |
| abstract_inverted_index.single-cell | 99 |
| abstract_inverted_index.substrates, | 34 |
| abstract_inverted_index.Perturbation | 139 |
| abstract_inverted_index.custom-built | 89 |
| abstract_inverted_index.environment, | 62 |
| abstract_inverted_index.hypothesized | 179 |
| abstract_inverted_index.perturbation | 190 |
| abstract_inverted_index.acidification. | 187, 209 |
| abstract_inverted_index.interleukin-10 | 126 |
| abstract_inverted_index.cytokine-regulated | 229 |
| abstract_inverted_index.LysoTracker-positive | 204 |
| abstract_inverted_index.necroptosis-cannibalism | 244 |
| cited_by_percentile_year | |
| countries_distinct_count | 1 |
| institutions_distinct_count | 12 |
| citation_normalized_percentile.value | 0.76989033 |
| citation_normalized_percentile.is_in_top_1_percent | False |
| citation_normalized_percentile.is_in_top_10_percent | False |