Genome‐wide investigation of a neuropathological cohort offers novel insights in the genetic risk underlying hallmark and co‐morbid lesions implicated in Alzheimer’s disease Article Swipe
 YOU?
·
· 2024
· Open Access
·
· DOI: https://doi.org/10.1002/alz.092254
YOU?
·
· 2024
· Open Access
·
· DOI: https://doi.org/10.1002/alz.092254
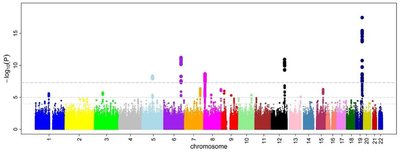
Background Alzheimer’s disease (AD) is a heterogenous disease with a strong heritability. Genetic studies are of irreplaceable value in elucidating the mechanisms that underly this disease. The classical genome‐wide association studies (GWAS) rely on ever‐increasing sample sizes and utilize clinical AD diagnosis to investigate genetic risk. Here, we aim to eliminate phenotypic heterogeneity by performing GWAS on a deeply phenotyped neuropathological cohort. Method Phenotypic data regarding hallmark and co‐morbid AD lesions was collected on 721 individuals. DNA was extracted from frozen or paraffin‐fixed cerebellar tissue. Subsequently, genetic data was generated using low‐coverage whole‐genome sequencing (lcWGS) followed by imputation based on 1000 Genomes Project reference panel using GLIMPSE. Linear or logistic regression was performed in PLINK correcting for age, gender and the first 3 PCs for every phenotype. Investigated phenotypes include but are not limited to: Amyloid phases, Braak‐NFT stage, cerebral amyloid angiopathy (CAA) severity and type, pTDP presence in CA1 and dentate gyrus, alpha‐synuclein spread/presence, age‐related tau‐astrogliopathy (ARTAG) and Hirano bodies. Result We discovered genome‐wide significant hits for well‐established AD lesions such as tau tangles, amyloid plaques and CAA. Furthermore, significant hits for co‐morbid lesions like alpha‐synuclein stages, granulovacuolar degeneration and Hirano bodies (extracellular actin aggregates in the CA1) were found. Suggestive associations were discovered for pTDP and ARTAG. Associated loci included proof‐of‐concept findings such as strong relations between amyloid and CAA with APOE SNPs, but also included several loci that have not been previously linked to the specific phenotype or AD in general. Conclusion Implementing phenotypic information on neuropathological lesions in a hypothesis‐free genome‐wide investigation as a preliminary screening rendered genome‐wide significant and suggestive loci which can contribute to a deeper understanding of the individual lesions observed in AD. From our preliminary results we observe the benefits of accurate phenotyping on GWAS power. Despite small sample size, we managed to detect genome‐wide significant associations for AD‐related lesions by eliminating phenotypic heterogeneity. Associated loci will be subjected to gene prioritization and functional validation analysis to determine the causal risk genes. For more commonly investigated hallmark lesions, replication analysis will be performed in publicly available cohorts.
Related Topics
- Type
- article
- Language
- en
- Landing Page
- https://doi.org/10.1002/alz.092254
- OA Status
- hybrid
- Related Works
- 10
- OpenAlex ID
- https://openalex.org/W4406050988
Raw OpenAlex JSON
- OpenAlex ID
-
https://openalex.org/W4406050988Canonical identifier for this work in OpenAlex
- DOI
-
https://doi.org/10.1002/alz.092254Digital Object Identifier
- Title
-
Genome‐wide investigation of a neuropathological cohort offers novel insights in the genetic risk underlying hallmark and co‐morbid lesions implicated in Alzheimer’s diseaseWork title
- Type
-
articleOpenAlex work type
- Language
-
enPrimary language
- Publication year
-
2024Year of publication
- Publication date
-
2024-12-01Full publication date if available
- Authors
-
Celeste Laureyssen, Klara Gawor, Jasper Van Dongen, Fahri Küçükali, Marta J. Koper, Alicja Ronisz, Markus Otto, Christine A. F. Von Arnim, Philip Van Damme, Rik Vandenberghe, Dietmar Rudolf Thal, Kristel SleegersList of authors in order
- Landing page
-
https://doi.org/10.1002/alz.092254Publisher landing page
- Open access
-
YesWhether a free full text is available
- OA status
-
hybridOpen access status per OpenAlex
- OA URL
-
https://doi.org/10.1002/alz.092254Direct OA link when available
- Concepts
-
Genome-wide association study, Single-nucleotide polymorphism, Biology, Disease, Cerebral amyloid angiopathy, Phenotype, Genetics, Alzheimer's disease, Missing heritability problem, Genetic association, Pathology, Medicine, Genotype, Gene, DementiaTop concepts (fields/topics) attached by OpenAlex
- Cited by
-
0Total citation count in OpenAlex
- Related works (count)
-
10Other works algorithmically related by OpenAlex
Full payload
| id | https://openalex.org/W4406050988 |
|---|---|
| doi | https://doi.org/10.1002/alz.092254 |
| ids.doi | https://doi.org/10.1002/alz.092254 |
| ids.pmid | https://pubmed.ncbi.nlm.nih.gov/39750869 |
| ids.openalex | https://openalex.org/W4406050988 |
| fwci | 0.0 |
| mesh[0].qualifier_ui | |
| mesh[0].descriptor_ui | D006801 |
| mesh[0].is_major_topic | False |
| mesh[0].qualifier_name | |
| mesh[0].descriptor_name | Humans |
| mesh[1].qualifier_ui | Q000235 |
| mesh[1].descriptor_ui | D000544 |
| mesh[1].is_major_topic | True |
| mesh[1].qualifier_name | genetics |
| mesh[1].descriptor_name | Alzheimer Disease |
| mesh[2].qualifier_ui | Q000473 |
| mesh[2].descriptor_ui | D000544 |
| mesh[2].is_major_topic | True |
| mesh[2].qualifier_name | pathology |
| mesh[2].descriptor_name | Alzheimer Disease |
| mesh[3].qualifier_ui | |
| mesh[3].descriptor_ui | D055106 |
| mesh[3].is_major_topic | True |
| mesh[3].qualifier_name | |
| mesh[3].descriptor_name | Genome-Wide Association Study |
| mesh[4].qualifier_ui | |
| mesh[4].descriptor_ui | D005260 |
| mesh[4].is_major_topic | False |
| mesh[4].qualifier_name | |
| mesh[4].descriptor_name | Female |
| mesh[5].qualifier_ui | |
| mesh[5].descriptor_ui | D008297 |
| mesh[5].is_major_topic | False |
| mesh[5].qualifier_name | |
| mesh[5].descriptor_name | Male |
| mesh[6].qualifier_ui | |
| mesh[6].descriptor_ui | D000368 |
| mesh[6].is_major_topic | False |
| mesh[6].qualifier_name | |
| mesh[6].descriptor_name | Aged |
| mesh[7].qualifier_ui | |
| mesh[7].descriptor_ui | D010641 |
| mesh[7].is_major_topic | False |
| mesh[7].qualifier_name | |
| mesh[7].descriptor_name | Phenotype |
| mesh[8].qualifier_ui | |
| mesh[8].descriptor_ui | D000369 |
| mesh[8].is_major_topic | False |
| mesh[8].qualifier_name | |
| mesh[8].descriptor_name | Aged, 80 and over |
| mesh[9].qualifier_ui | |
| mesh[9].descriptor_ui | D015331 |
| mesh[9].is_major_topic | False |
| mesh[9].qualifier_name | |
| mesh[9].descriptor_name | Cohort Studies |
| mesh[10].qualifier_ui | Q000235 |
| mesh[10].descriptor_ui | D016657 |
| mesh[10].is_major_topic | False |
| mesh[10].qualifier_name | genetics |
| mesh[10].descriptor_name | Cerebral Amyloid Angiopathy |
| mesh[11].qualifier_ui | Q000473 |
| mesh[11].descriptor_ui | D016657 |
| mesh[11].is_major_topic | False |
| mesh[11].qualifier_name | pathology |
| mesh[11].descriptor_name | Cerebral Amyloid Angiopathy |
| mesh[12].qualifier_ui | Q000473 |
| mesh[12].descriptor_ui | D001921 |
| mesh[12].is_major_topic | False |
| mesh[12].qualifier_name | pathology |
| mesh[12].descriptor_name | Brain |
| mesh[13].qualifier_ui | |
| mesh[13].descriptor_ui | D020022 |
| mesh[13].is_major_topic | False |
| mesh[13].qualifier_name | |
| mesh[13].descriptor_name | Genetic Predisposition to Disease |
| mesh[14].qualifier_ui | |
| mesh[14].descriptor_ui | D006801 |
| mesh[14].is_major_topic | False |
| mesh[14].qualifier_name | |
| mesh[14].descriptor_name | Humans |
| mesh[15].qualifier_ui | Q000235 |
| mesh[15].descriptor_ui | D000544 |
| mesh[15].is_major_topic | True |
| mesh[15].qualifier_name | genetics |
| mesh[15].descriptor_name | Alzheimer Disease |
| mesh[16].qualifier_ui | Q000473 |
| mesh[16].descriptor_ui | D000544 |
| mesh[16].is_major_topic | True |
| mesh[16].qualifier_name | pathology |
| mesh[16].descriptor_name | Alzheimer Disease |
| mesh[17].qualifier_ui | |
| mesh[17].descriptor_ui | D055106 |
| mesh[17].is_major_topic | True |
| mesh[17].qualifier_name | |
| mesh[17].descriptor_name | Genome-Wide Association Study |
| mesh[18].qualifier_ui | |
| mesh[18].descriptor_ui | D005260 |
| mesh[18].is_major_topic | False |
| mesh[18].qualifier_name | |
| mesh[18].descriptor_name | Female |
| mesh[19].qualifier_ui | |
| mesh[19].descriptor_ui | D008297 |
| mesh[19].is_major_topic | False |
| mesh[19].qualifier_name | |
| mesh[19].descriptor_name | Male |
| mesh[20].qualifier_ui | |
| mesh[20].descriptor_ui | D000368 |
| mesh[20].is_major_topic | False |
| mesh[20].qualifier_name | |
| mesh[20].descriptor_name | Aged |
| mesh[21].qualifier_ui | |
| mesh[21].descriptor_ui | D010641 |
| mesh[21].is_major_topic | False |
| mesh[21].qualifier_name | |
| mesh[21].descriptor_name | Phenotype |
| mesh[22].qualifier_ui | |
| mesh[22].descriptor_ui | D000369 |
| mesh[22].is_major_topic | False |
| mesh[22].qualifier_name | |
| mesh[22].descriptor_name | Aged, 80 and over |
| mesh[23].qualifier_ui | |
| mesh[23].descriptor_ui | D015331 |
| mesh[23].is_major_topic | False |
| mesh[23].qualifier_name | |
| mesh[23].descriptor_name | Cohort Studies |
| mesh[24].qualifier_ui | Q000235 |
| mesh[24].descriptor_ui | D016657 |
| mesh[24].is_major_topic | False |
| mesh[24].qualifier_name | genetics |
| mesh[24].descriptor_name | Cerebral Amyloid Angiopathy |
| mesh[25].qualifier_ui | Q000473 |
| mesh[25].descriptor_ui | D016657 |
| mesh[25].is_major_topic | False |
| mesh[25].qualifier_name | pathology |
| mesh[25].descriptor_name | Cerebral Amyloid Angiopathy |
| mesh[26].qualifier_ui | Q000473 |
| mesh[26].descriptor_ui | D001921 |
| mesh[26].is_major_topic | False |
| mesh[26].qualifier_name | pathology |
| mesh[26].descriptor_name | Brain |
| mesh[27].qualifier_ui | |
| mesh[27].descriptor_ui | D020022 |
| mesh[27].is_major_topic | False |
| mesh[27].qualifier_name | |
| mesh[27].descriptor_name | Genetic Predisposition to Disease |
| type | article |
| title | Genome‐wide investigation of a neuropathological cohort offers novel insights in the genetic risk underlying hallmark and co‐morbid lesions implicated in Alzheimer’s disease |
| biblio.issue | S1 |
| biblio.volume | 20 |
| biblio.last_page | e092254 |
| biblio.first_page | e092254 |
| topics[0].id | https://openalex.org/T10086 |
| topics[0].field.id | https://openalex.org/fields/27 |
| topics[0].field.display_name | Medicine |
| topics[0].score | 0.9983000159263611 |
| topics[0].domain.id | https://openalex.org/domains/4 |
| topics[0].domain.display_name | Health Sciences |
| topics[0].subfield.id | https://openalex.org/subfields/2737 |
| topics[0].subfield.display_name | Physiology |
| topics[0].display_name | Alzheimer's disease research and treatments |
| topics[1].id | https://openalex.org/T10009 |
| topics[1].field.id | https://openalex.org/fields/27 |
| topics[1].field.display_name | Medicine |
| topics[1].score | 0.9887999892234802 |
| topics[1].domain.id | https://openalex.org/domains/4 |
| topics[1].domain.display_name | Health Sciences |
| topics[1].subfield.id | https://openalex.org/subfields/2738 |
| topics[1].subfield.display_name | Psychiatry and Mental health |
| topics[1].display_name | Dementia and Cognitive Impairment Research |
| topics[2].id | https://openalex.org/T10261 |
| topics[2].field.id | https://openalex.org/fields/13 |
| topics[2].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[2].score | 0.9858999848365784 |
| topics[2].domain.id | https://openalex.org/domains/1 |
| topics[2].domain.display_name | Life Sciences |
| topics[2].subfield.id | https://openalex.org/subfields/1311 |
| topics[2].subfield.display_name | Genetics |
| topics[2].display_name | Genetic Associations and Epidemiology |
| is_xpac | False |
| apc_list.value | 4000 |
| apc_list.currency | USD |
| apc_list.value_usd | 4000 |
| apc_paid.value | 4000 |
| apc_paid.currency | USD |
| apc_paid.value_usd | 4000 |
| concepts[0].id | https://openalex.org/C106208931 |
| concepts[0].level | 5 |
| concepts[0].score | 0.7672463655471802 |
| concepts[0].wikidata | https://www.wikidata.org/wiki/Q1098876 |
| concepts[0].display_name | Genome-wide association study |
| concepts[1].id | https://openalex.org/C153209595 |
| concepts[1].level | 4 |
| concepts[1].score | 0.5855335593223572 |
| concepts[1].wikidata | https://www.wikidata.org/wiki/Q501128 |
| concepts[1].display_name | Single-nucleotide polymorphism |
| concepts[2].id | https://openalex.org/C86803240 |
| concepts[2].level | 0 |
| concepts[2].score | 0.561626672744751 |
| concepts[2].wikidata | https://www.wikidata.org/wiki/Q420 |
| concepts[2].display_name | Biology |
| concepts[3].id | https://openalex.org/C2779134260 |
| concepts[3].level | 2 |
| concepts[3].score | 0.5128824710845947 |
| concepts[3].wikidata | https://www.wikidata.org/wiki/Q12136 |
| concepts[3].display_name | Disease |
| concepts[4].id | https://openalex.org/C2777790613 |
| concepts[4].level | 4 |
| concepts[4].score | 0.507850706577301 |
| concepts[4].wikidata | https://www.wikidata.org/wiki/Q191562 |
| concepts[4].display_name | Cerebral amyloid angiopathy |
| concepts[5].id | https://openalex.org/C127716648 |
| concepts[5].level | 3 |
| concepts[5].score | 0.4828489422798157 |
| concepts[5].wikidata | https://www.wikidata.org/wiki/Q104053 |
| concepts[5].display_name | Phenotype |
| concepts[6].id | https://openalex.org/C54355233 |
| concepts[6].level | 1 |
| concepts[6].score | 0.47722509503364563 |
| concepts[6].wikidata | https://www.wikidata.org/wiki/Q7162 |
| concepts[6].display_name | Genetics |
| concepts[7].id | https://openalex.org/C502032728 |
| concepts[7].level | 3 |
| concepts[7].score | 0.4657832980155945 |
| concepts[7].wikidata | https://www.wikidata.org/wiki/Q11081 |
| concepts[7].display_name | Alzheimer's disease |
| concepts[8].id | https://openalex.org/C144621757 |
| concepts[8].level | 5 |
| concepts[8].score | 0.44207707047462463 |
| concepts[8].wikidata | https://www.wikidata.org/wiki/Q3083720 |
| concepts[8].display_name | Missing heritability problem |
| concepts[9].id | https://openalex.org/C186413461 |
| concepts[9].level | 5 |
| concepts[9].score | 0.41865700483322144 |
| concepts[9].wikidata | https://www.wikidata.org/wiki/Q744727 |
| concepts[9].display_name | Genetic association |
| concepts[10].id | https://openalex.org/C142724271 |
| concepts[10].level | 1 |
| concepts[10].score | 0.38222163915634155 |
| concepts[10].wikidata | https://www.wikidata.org/wiki/Q7208 |
| concepts[10].display_name | Pathology |
| concepts[11].id | https://openalex.org/C71924100 |
| concepts[11].level | 0 |
| concepts[11].score | 0.23652005195617676 |
| concepts[11].wikidata | https://www.wikidata.org/wiki/Q11190 |
| concepts[11].display_name | Medicine |
| concepts[12].id | https://openalex.org/C135763542 |
| concepts[12].level | 3 |
| concepts[12].score | 0.19743022322654724 |
| concepts[12].wikidata | https://www.wikidata.org/wiki/Q106016 |
| concepts[12].display_name | Genotype |
| concepts[13].id | https://openalex.org/C104317684 |
| concepts[13].level | 2 |
| concepts[13].score | 0.19230356812477112 |
| concepts[13].wikidata | https://www.wikidata.org/wiki/Q7187 |
| concepts[13].display_name | Gene |
| concepts[14].id | https://openalex.org/C2779483572 |
| concepts[14].level | 3 |
| concepts[14].score | 0.17272180318832397 |
| concepts[14].wikidata | https://www.wikidata.org/wiki/Q83030 |
| concepts[14].display_name | Dementia |
| keywords[0].id | https://openalex.org/keywords/genome-wide-association-study |
| keywords[0].score | 0.7672463655471802 |
| keywords[0].display_name | Genome-wide association study |
| keywords[1].id | https://openalex.org/keywords/single-nucleotide-polymorphism |
| keywords[1].score | 0.5855335593223572 |
| keywords[1].display_name | Single-nucleotide polymorphism |
| keywords[2].id | https://openalex.org/keywords/biology |
| keywords[2].score | 0.561626672744751 |
| keywords[2].display_name | Biology |
| keywords[3].id | https://openalex.org/keywords/disease |
| keywords[3].score | 0.5128824710845947 |
| keywords[3].display_name | Disease |
| keywords[4].id | https://openalex.org/keywords/cerebral-amyloid-angiopathy |
| keywords[4].score | 0.507850706577301 |
| keywords[4].display_name | Cerebral amyloid angiopathy |
| keywords[5].id | https://openalex.org/keywords/phenotype |
| keywords[5].score | 0.4828489422798157 |
| keywords[5].display_name | Phenotype |
| keywords[6].id | https://openalex.org/keywords/genetics |
| keywords[6].score | 0.47722509503364563 |
| keywords[6].display_name | Genetics |
| keywords[7].id | https://openalex.org/keywords/alzheimers-disease |
| keywords[7].score | 0.4657832980155945 |
| keywords[7].display_name | Alzheimer's disease |
| keywords[8].id | https://openalex.org/keywords/missing-heritability-problem |
| keywords[8].score | 0.44207707047462463 |
| keywords[8].display_name | Missing heritability problem |
| keywords[9].id | https://openalex.org/keywords/genetic-association |
| keywords[9].score | 0.41865700483322144 |
| keywords[9].display_name | Genetic association |
| keywords[10].id | https://openalex.org/keywords/pathology |
| keywords[10].score | 0.38222163915634155 |
| keywords[10].display_name | Pathology |
| keywords[11].id | https://openalex.org/keywords/medicine |
| keywords[11].score | 0.23652005195617676 |
| keywords[11].display_name | Medicine |
| keywords[12].id | https://openalex.org/keywords/genotype |
| keywords[12].score | 0.19743022322654724 |
| keywords[12].display_name | Genotype |
| keywords[13].id | https://openalex.org/keywords/gene |
| keywords[13].score | 0.19230356812477112 |
| keywords[13].display_name | Gene |
| keywords[14].id | https://openalex.org/keywords/dementia |
| keywords[14].score | 0.17272180318832397 |
| keywords[14].display_name | Dementia |
| language | en |
| locations[0].id | doi:10.1002/alz.092254 |
| locations[0].is_oa | True |
| locations[0].source.id | https://openalex.org/S108427512 |
| locations[0].source.issn | 1552-5260, 1552-5279 |
| locations[0].source.type | journal |
| locations[0].source.is_oa | False |
| locations[0].source.issn_l | 1552-5260 |
| locations[0].source.is_core | True |
| locations[0].source.is_in_doaj | False |
| locations[0].source.display_name | Alzheimer s & Dementia |
| locations[0].source.host_organization | https://openalex.org/P4310320595 |
| locations[0].source.host_organization_name | Wiley |
| locations[0].source.host_organization_lineage | https://openalex.org/P4310320595 |
| locations[0].license | cc-by |
| locations[0].pdf_url | |
| locations[0].version | publishedVersion |
| locations[0].raw_type | journal-article |
| locations[0].license_id | https://openalex.org/licenses/cc-by |
| locations[0].is_accepted | True |
| locations[0].is_published | True |
| locations[0].raw_source_name | Alzheimer's & Dementia |
| locations[0].landing_page_url | https://doi.org/10.1002/alz.092254 |
| locations[1].id | pmid:39750869 |
| locations[1].is_oa | False |
| locations[1].source.id | https://openalex.org/S4306525036 |
| locations[1].source.issn | |
| locations[1].source.type | repository |
| locations[1].source.is_oa | False |
| locations[1].source.issn_l | |
| locations[1].source.is_core | False |
| locations[1].source.is_in_doaj | False |
| locations[1].source.display_name | PubMed |
| locations[1].source.host_organization | https://openalex.org/I1299303238 |
| locations[1].source.host_organization_name | National Institutes of Health |
| locations[1].source.host_organization_lineage | https://openalex.org/I1299303238 |
| locations[1].license | |
| locations[1].pdf_url | |
| locations[1].version | publishedVersion |
| locations[1].raw_type | |
| locations[1].license_id | |
| locations[1].is_accepted | True |
| locations[1].is_published | True |
| locations[1].raw_source_name | Alzheimer's & dementia : the journal of the Alzheimer's Association |
| locations[1].landing_page_url | https://pubmed.ncbi.nlm.nih.gov/39750869 |
| locations[2].id | pmh:oai:lirias2repo.kuleuven.be:20.500.12942/756998 |
| locations[2].is_oa | True |
| locations[2].source.id | https://openalex.org/S4306401954 |
| locations[2].source.issn | |
| locations[2].source.type | repository |
| locations[2].source.is_oa | False |
| locations[2].source.issn_l | |
| locations[2].source.is_core | False |
| locations[2].source.is_in_doaj | False |
| locations[2].source.display_name | Lirias (KU Leuven) |
| locations[2].source.host_organization | https://openalex.org/I99464096 |
| locations[2].source.host_organization_name | KU Leuven |
| locations[2].source.host_organization_lineage | https://openalex.org/I99464096 |
| locations[2].license | cc-by |
| locations[2].pdf_url | |
| locations[2].version | acceptedVersion |
| locations[2].raw_type | info:eu-repo/semantics/lecture |
| locations[2].license_id | https://openalex.org/licenses/cc-by |
| locations[2].is_accepted | True |
| locations[2].is_published | False |
| locations[2].raw_source_name | |
| locations[2].landing_page_url | https://lirias.kuleuven.be/handle/20.500.12942/756998 |
| locations[3].id | pmh:oai:pubmedcentral.nih.gov:11709958 |
| locations[3].is_oa | True |
| locations[3].source.id | https://openalex.org/S2764455111 |
| locations[3].source.issn | |
| locations[3].source.type | repository |
| locations[3].source.is_oa | False |
| locations[3].source.issn_l | |
| locations[3].source.is_core | False |
| locations[3].source.is_in_doaj | False |
| locations[3].source.display_name | PubMed Central |
| locations[3].source.host_organization | https://openalex.org/I1299303238 |
| locations[3].source.host_organization_name | National Institutes of Health |
| locations[3].source.host_organization_lineage | https://openalex.org/I1299303238 |
| locations[3].license | other-oa |
| locations[3].pdf_url | |
| locations[3].version | submittedVersion |
| locations[3].raw_type | Text |
| locations[3].license_id | https://openalex.org/licenses/other-oa |
| locations[3].is_accepted | False |
| locations[3].is_published | False |
| locations[3].raw_source_name | Alzheimers Dement |
| locations[3].landing_page_url | https://www.ncbi.nlm.nih.gov/pmc/articles/11709958 |
| indexed_in | crossref, pubmed |
| authorships[0].author.id | https://openalex.org/A5092973172 |
| authorships[0].author.orcid | https://orcid.org/0000-0002-7373-2099 |
| authorships[0].author.display_name | Celeste Laureyssen |
| authorships[0].countries | BE |
| authorships[0].affiliations[0].institution_ids | https://openalex.org/I149213910 |
| authorships[0].affiliations[0].raw_affiliation_string | Department of Biomedical Sciences, University of Antwerp, Antwerp Belgium |
| authorships[0].affiliations[1].institution_ids | https://openalex.org/I4210132484 |
| authorships[0].affiliations[1].raw_affiliation_string | Complex Genetics of Alzheimer’s Disease Group, VIB Center for Molecular Neurology, VIB, Antwerp Belgium |
| authorships[0].institutions[0].id | https://openalex.org/I149213910 |
| authorships[0].institutions[0].ror | https://ror.org/008x57b05 |
| authorships[0].institutions[0].type | education |
| authorships[0].institutions[0].lineage | https://openalex.org/I149213910 |
| authorships[0].institutions[0].country_code | BE |
| authorships[0].institutions[0].display_name | University of Antwerp |
| authorships[0].institutions[1].id | https://openalex.org/I4210132484 |
| authorships[0].institutions[1].ror | https://ror.org/041x7eh14 |
| authorships[0].institutions[1].type | facility |
| authorships[0].institutions[1].lineage | https://openalex.org/I149213910, https://openalex.org/I2802017950, https://openalex.org/I4210132484 |
| authorships[0].institutions[1].country_code | BE |
| authorships[0].institutions[1].display_name | VIB-UAntwerp Center for Molecular Neurology |
| authorships[0].author_position | first |
| authorships[0].raw_author_name | Celeste Laureyssen |
| authorships[0].is_corresponding | False |
| authorships[0].raw_affiliation_strings | Complex Genetics of Alzheimer’s Disease Group, VIB Center for Molecular Neurology, VIB, Antwerp Belgium, Department of Biomedical Sciences, University of Antwerp, Antwerp Belgium |
| authorships[1].author.id | https://openalex.org/A5015248328 |
| authorships[1].author.orcid | https://orcid.org/0000-0001-7571-5420 |
| authorships[1].author.display_name | Klara Gawor |
| authorships[1].countries | BE |
| authorships[1].affiliations[0].institution_ids | https://openalex.org/I99464096 |
| authorships[1].affiliations[0].raw_affiliation_string | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[1].institutions[0].id | https://openalex.org/I99464096 |
| authorships[1].institutions[0].ror | https://ror.org/05f950310 |
| authorships[1].institutions[0].type | education |
| authorships[1].institutions[0].lineage | https://openalex.org/I99464096 |
| authorships[1].institutions[0].country_code | BE |
| authorships[1].institutions[0].display_name | KU Leuven |
| authorships[1].author_position | middle |
| authorships[1].raw_author_name | Klara Gawor |
| authorships[1].is_corresponding | False |
| authorships[1].raw_affiliation_strings | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[2].author.id | https://openalex.org/A5081371190 |
| authorships[2].author.orcid | https://orcid.org/0000-0001-5599-0046 |
| authorships[2].author.display_name | Jasper Van Dongen |
| authorships[2].countries | BE |
| authorships[2].affiliations[0].institution_ids | https://openalex.org/I4210132484 |
| authorships[2].affiliations[0].raw_affiliation_string | Complex Genetics of Alzheimer’s Disease group, VIB‐UAntwerp Center for Molecular Neurology, Antwerp Belgium |
| authorships[2].affiliations[1].institution_ids | https://openalex.org/I149213910 |
| authorships[2].affiliations[1].raw_affiliation_string | University of Antwerp, Antwerp Belgium |
| authorships[2].institutions[0].id | https://openalex.org/I149213910 |
| authorships[2].institutions[0].ror | https://ror.org/008x57b05 |
| authorships[2].institutions[0].type | education |
| authorships[2].institutions[0].lineage | https://openalex.org/I149213910 |
| authorships[2].institutions[0].country_code | BE |
| authorships[2].institutions[0].display_name | University of Antwerp |
| authorships[2].institutions[1].id | https://openalex.org/I4210132484 |
| authorships[2].institutions[1].ror | https://ror.org/041x7eh14 |
| authorships[2].institutions[1].type | facility |
| authorships[2].institutions[1].lineage | https://openalex.org/I149213910, https://openalex.org/I2802017950, https://openalex.org/I4210132484 |
| authorships[2].institutions[1].country_code | BE |
| authorships[2].institutions[1].display_name | VIB-UAntwerp Center for Molecular Neurology |
| authorships[2].author_position | middle |
| authorships[2].raw_author_name | Jasper Van Dongen |
| authorships[2].is_corresponding | False |
| authorships[2].raw_affiliation_strings | Complex Genetics of Alzheimer’s Disease group, VIB‐UAntwerp Center for Molecular Neurology, Antwerp Belgium, University of Antwerp, Antwerp Belgium |
| authorships[3].author.id | https://openalex.org/A5056524762 |
| authorships[3].author.orcid | https://orcid.org/0000-0002-3835-9639 |
| authorships[3].author.display_name | Fahri Küçükali |
| authorships[3].countries | BE |
| authorships[3].affiliations[0].institution_ids | https://openalex.org/I149213910 |
| authorships[3].affiliations[0].raw_affiliation_string | Department of Biomedical Sciences, University of Antwerp, Antwerp Belgium |
| authorships[3].affiliations[1].institution_ids | https://openalex.org/I4210132484 |
| authorships[3].affiliations[1].raw_affiliation_string | Complex Genetics of Alzheimer’s Disease Group, VIB Center for Molecular Neurology, VIB, Antwerp Belgium |
| authorships[3].institutions[0].id | https://openalex.org/I149213910 |
| authorships[3].institutions[0].ror | https://ror.org/008x57b05 |
| authorships[3].institutions[0].type | education |
| authorships[3].institutions[0].lineage | https://openalex.org/I149213910 |
| authorships[3].institutions[0].country_code | BE |
| authorships[3].institutions[0].display_name | University of Antwerp |
| authorships[3].institutions[1].id | https://openalex.org/I4210132484 |
| authorships[3].institutions[1].ror | https://ror.org/041x7eh14 |
| authorships[3].institutions[1].type | facility |
| authorships[3].institutions[1].lineage | https://openalex.org/I149213910, https://openalex.org/I2802017950, https://openalex.org/I4210132484 |
| authorships[3].institutions[1].country_code | BE |
| authorships[3].institutions[1].display_name | VIB-UAntwerp Center for Molecular Neurology |
| authorships[3].author_position | middle |
| authorships[3].raw_author_name | Fahri Küçükali |
| authorships[3].is_corresponding | False |
| authorships[3].raw_affiliation_strings | Complex Genetics of Alzheimer’s Disease Group, VIB Center for Molecular Neurology, VIB, Antwerp Belgium, Department of Biomedical Sciences, University of Antwerp, Antwerp Belgium |
| authorships[4].author.id | https://openalex.org/A5031573702 |
| authorships[4].author.orcid | https://orcid.org/0000-0003-2255-6088 |
| authorships[4].author.display_name | Marta J. Koper |
| authorships[4].countries | BE |
| authorships[4].affiliations[0].institution_ids | https://openalex.org/I99464096 |
| authorships[4].affiliations[0].raw_affiliation_string | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[4].institutions[0].id | https://openalex.org/I99464096 |
| authorships[4].institutions[0].ror | https://ror.org/05f950310 |
| authorships[4].institutions[0].type | education |
| authorships[4].institutions[0].lineage | https://openalex.org/I99464096 |
| authorships[4].institutions[0].country_code | BE |
| authorships[4].institutions[0].display_name | KU Leuven |
| authorships[4].author_position | middle |
| authorships[4].raw_author_name | Marta J Koper |
| authorships[4].is_corresponding | False |
| authorships[4].raw_affiliation_strings | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[5].author.id | https://openalex.org/A5009935278 |
| authorships[5].author.orcid | https://orcid.org/0000-0002-4310-0445 |
| authorships[5].author.display_name | Alicja Ronisz |
| authorships[5].countries | BE |
| authorships[5].affiliations[0].institution_ids | https://openalex.org/I99464096 |
| authorships[5].affiliations[0].raw_affiliation_string | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[5].institutions[0].id | https://openalex.org/I99464096 |
| authorships[5].institutions[0].ror | https://ror.org/05f950310 |
| authorships[5].institutions[0].type | education |
| authorships[5].institutions[0].lineage | https://openalex.org/I99464096 |
| authorships[5].institutions[0].country_code | BE |
| authorships[5].institutions[0].display_name | KU Leuven |
| authorships[5].author_position | middle |
| authorships[5].raw_author_name | Alicja Ronisz |
| authorships[5].is_corresponding | False |
| authorships[5].raw_affiliation_strings | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[6].author.id | https://openalex.org/A5071308068 |
| authorships[6].author.orcid | https://orcid.org/0000-0003-4273-4267 |
| authorships[6].author.display_name | Markus Otto |
| authorships[6].countries | DE |
| authorships[6].affiliations[0].institution_ids | https://openalex.org/I4210155144 |
| authorships[6].affiliations[0].raw_affiliation_string | Department of Neurology, Ulm University Hospital, Ulm Germany |
| authorships[6].institutions[0].id | https://openalex.org/I4210155144 |
| authorships[6].institutions[0].ror | https://ror.org/05emabm63 |
| authorships[6].institutions[0].type | healthcare |
| authorships[6].institutions[0].lineage | https://openalex.org/I4210155144 |
| authorships[6].institutions[0].country_code | DE |
| authorships[6].institutions[0].display_name | University Hospital Ulm |
| authorships[6].author_position | middle |
| authorships[6].raw_author_name | Markus Otto |
| authorships[6].is_corresponding | False |
| authorships[6].raw_affiliation_strings | Department of Neurology, Ulm University Hospital, Ulm Germany |
| authorships[7].author.id | https://openalex.org/A5080821885 |
| authorships[7].author.orcid | https://orcid.org/0000-0002-8614-2223 |
| authorships[7].author.display_name | Christine A. F. Von Arnim |
| authorships[7].countries | DE |
| authorships[7].affiliations[0].institution_ids | https://openalex.org/I4210116730 |
| authorships[7].affiliations[0].raw_affiliation_string | University Medical Center Göttingen, Göttingen Germany |
| authorships[7].institutions[0].id | https://openalex.org/I4210116730 |
| authorships[7].institutions[0].ror | https://ror.org/021ft0n22 |
| authorships[7].institutions[0].type | healthcare |
| authorships[7].institutions[0].lineage | https://openalex.org/I4210116730 |
| authorships[7].institutions[0].country_code | DE |
| authorships[7].institutions[0].display_name | Universitätsmedizin Göttingen |
| authorships[7].author_position | middle |
| authorships[7].raw_author_name | Christine von Arnim |
| authorships[7].is_corresponding | False |
| authorships[7].raw_affiliation_strings | University Medical Center Göttingen, Göttingen Germany |
| authorships[8].author.id | https://openalex.org/A5013023361 |
| authorships[8].author.orcid | https://orcid.org/0000-0002-4010-2357 |
| authorships[8].author.display_name | Philip Van Damme |
| authorships[8].countries | BE |
| authorships[8].affiliations[0].institution_ids | https://openalex.org/I99464096 |
| authorships[8].affiliations[0].raw_affiliation_string | Department of Neurosciences, KU Leuven, Leuven Belgium |
| authorships[8].affiliations[1].institution_ids | https://openalex.org/I2800231306 |
| authorships[8].affiliations[1].raw_affiliation_string | UZ Leuven, Leuven Belgium |
| authorships[8].institutions[0].id | https://openalex.org/I99464096 |
| authorships[8].institutions[0].ror | https://ror.org/05f950310 |
| authorships[8].institutions[0].type | education |
| authorships[8].institutions[0].lineage | https://openalex.org/I99464096 |
| authorships[8].institutions[0].country_code | BE |
| authorships[8].institutions[0].display_name | KU Leuven |
| authorships[8].institutions[1].id | https://openalex.org/I2800231306 |
| authorships[8].institutions[1].ror | https://ror.org/0424bsv16 |
| authorships[8].institutions[1].type | healthcare |
| authorships[8].institutions[1].lineage | https://openalex.org/I2800231306 |
| authorships[8].institutions[1].country_code | BE |
| authorships[8].institutions[1].display_name | Universitair Ziekenhuis Leuven |
| authorships[8].author_position | middle |
| authorships[8].raw_author_name | Philip Van Damme |
| authorships[8].is_corresponding | False |
| authorships[8].raw_affiliation_strings | Department of Neurosciences, KU Leuven, Leuven Belgium, UZ Leuven, Leuven Belgium |
| authorships[9].author.id | https://openalex.org/A5071341444 |
| authorships[9].author.orcid | https://orcid.org/0000-0001-6237-2502 |
| authorships[9].author.display_name | Rik Vandenberghe |
| authorships[9].countries | BE |
| authorships[9].affiliations[0].institution_ids | https://openalex.org/I4210153928 |
| authorships[9].affiliations[0].raw_affiliation_string | Alzheimer Research Centre KU Leuven, Leuven Brain Institute, Leuven Belgium |
| authorships[9].affiliations[1].institution_ids | https://openalex.org/I99464096 |
| authorships[9].affiliations[1].raw_affiliation_string | KU Leuven, Leuven Belgium |
| authorships[9].institutions[0].id | https://openalex.org/I99464096 |
| authorships[9].institutions[0].ror | https://ror.org/05f950310 |
| authorships[9].institutions[0].type | education |
| authorships[9].institutions[0].lineage | https://openalex.org/I99464096 |
| authorships[9].institutions[0].country_code | BE |
| authorships[9].institutions[0].display_name | KU Leuven |
| authorships[9].institutions[1].id | https://openalex.org/I4210153928 |
| authorships[9].institutions[1].ror | https://ror.org/045c7t348 |
| authorships[9].institutions[1].type | facility |
| authorships[9].institutions[1].lineage | https://openalex.org/I2802017950, https://openalex.org/I4210153928, https://openalex.org/I99464096 |
| authorships[9].institutions[1].country_code | BE |
| authorships[9].institutions[1].display_name | VIB-KU Leuven Center for Brain & Disease Research |
| authorships[9].author_position | middle |
| authorships[9].raw_author_name | Rik Vandenberghe |
| authorships[9].is_corresponding | False |
| authorships[9].raw_affiliation_strings | Alzheimer Research Centre KU Leuven, Leuven Brain Institute, Leuven Belgium, KU Leuven, Leuven Belgium |
| authorships[10].author.id | https://openalex.org/A5023861108 |
| authorships[10].author.orcid | https://orcid.org/0000-0002-1036-1075 |
| authorships[10].author.display_name | Dietmar Rudolf Thal |
| authorships[10].countries | BE |
| authorships[10].affiliations[0].institution_ids | https://openalex.org/I2800231306 |
| authorships[10].affiliations[0].raw_affiliation_string | UZ Leuven, Leuven Belgium |
| authorships[10].affiliations[1].institution_ids | https://openalex.org/I99464096 |
| authorships[10].affiliations[1].raw_affiliation_string | Laboratory for Neuropathology, KU Leuven, Leuven Belgium |
| authorships[10].institutions[0].id | https://openalex.org/I99464096 |
| authorships[10].institutions[0].ror | https://ror.org/05f950310 |
| authorships[10].institutions[0].type | education |
| authorships[10].institutions[0].lineage | https://openalex.org/I99464096 |
| authorships[10].institutions[0].country_code | BE |
| authorships[10].institutions[0].display_name | KU Leuven |
| authorships[10].institutions[1].id | https://openalex.org/I2800231306 |
| authorships[10].institutions[1].ror | https://ror.org/0424bsv16 |
| authorships[10].institutions[1].type | healthcare |
| authorships[10].institutions[1].lineage | https://openalex.org/I2800231306 |
| authorships[10].institutions[1].country_code | BE |
| authorships[10].institutions[1].display_name | Universitair Ziekenhuis Leuven |
| authorships[10].author_position | middle |
| authorships[10].raw_author_name | Dietmar Rudolf Thal |
| authorships[10].is_corresponding | False |
| authorships[10].raw_affiliation_strings | Laboratory for Neuropathology, KU Leuven, Leuven Belgium, UZ Leuven, Leuven Belgium |
| authorships[11].author.id | https://openalex.org/A5015407411 |
| authorships[11].author.orcid | https://orcid.org/0000-0002-0283-2332 |
| authorships[11].author.display_name | Kristel Sleegers |
| authorships[11].countries | BE |
| authorships[11].affiliations[0].institution_ids | https://openalex.org/I149213910 |
| authorships[11].affiliations[0].raw_affiliation_string | Department of Biomedical Sciences, University of Antwerp, Antwerp Belgium |
| authorships[11].affiliations[1].institution_ids | https://openalex.org/I4210132484 |
| authorships[11].affiliations[1].raw_affiliation_string | Complex Genetics of Alzheimer’s Disease group, VIB‐UAntwerp Center for Molecular Neurology, Antwerp Belgium |
| authorships[11].institutions[0].id | https://openalex.org/I149213910 |
| authorships[11].institutions[0].ror | https://ror.org/008x57b05 |
| authorships[11].institutions[0].type | education |
| authorships[11].institutions[0].lineage | https://openalex.org/I149213910 |
| authorships[11].institutions[0].country_code | BE |
| authorships[11].institutions[0].display_name | University of Antwerp |
| authorships[11].institutions[1].id | https://openalex.org/I4210132484 |
| authorships[11].institutions[1].ror | https://ror.org/041x7eh14 |
| authorships[11].institutions[1].type | facility |
| authorships[11].institutions[1].lineage | https://openalex.org/I149213910, https://openalex.org/I2802017950, https://openalex.org/I4210132484 |
| authorships[11].institutions[1].country_code | BE |
| authorships[11].institutions[1].display_name | VIB-UAntwerp Center for Molecular Neurology |
| authorships[11].author_position | last |
| authorships[11].raw_author_name | Kristel Sleegers |
| authorships[11].is_corresponding | False |
| authorships[11].raw_affiliation_strings | Complex Genetics of Alzheimer’s Disease group, VIB‐UAntwerp Center for Molecular Neurology, Antwerp Belgium, Department of Biomedical Sciences, University of Antwerp, Antwerp Belgium |
| has_content.pdf | False |
| has_content.grobid_xml | False |
| is_paratext | False |
| open_access.is_oa | True |
| open_access.oa_url | https://doi.org/10.1002/alz.092254 |
| open_access.oa_status | hybrid |
| open_access.any_repository_has_fulltext | False |
| created_date | 2025-01-04T00:00:00 |
| display_name | Genome‐wide investigation of a neuropathological cohort offers novel insights in the genetic risk underlying hallmark and co‐morbid lesions implicated in Alzheimer’s disease |
| has_fulltext | False |
| is_retracted | False |
| updated_date | 2025-11-06T03:46:38.306776 |
| primary_topic.id | https://openalex.org/T10086 |
| primary_topic.field.id | https://openalex.org/fields/27 |
| primary_topic.field.display_name | Medicine |
| primary_topic.score | 0.9983000159263611 |
| primary_topic.domain.id | https://openalex.org/domains/4 |
| primary_topic.domain.display_name | Health Sciences |
| primary_topic.subfield.id | https://openalex.org/subfields/2737 |
| primary_topic.subfield.display_name | Physiology |
| primary_topic.display_name | Alzheimer's disease research and treatments |
| related_works | https://openalex.org/W1884571866, https://openalex.org/W2409016390, https://openalex.org/W2160362734, https://openalex.org/W159064268, https://openalex.org/W2111896105, https://openalex.org/W2170506381, https://openalex.org/W1987620105, https://openalex.org/W1847429134, https://openalex.org/W2144000849, https://openalex.org/W2067245483 |
| cited_by_count | 0 |
| locations_count | 4 |
| best_oa_location.id | doi:10.1002/alz.092254 |
| best_oa_location.is_oa | True |
| best_oa_location.source.id | https://openalex.org/S108427512 |
| best_oa_location.source.issn | 1552-5260, 1552-5279 |
| best_oa_location.source.type | journal |
| best_oa_location.source.is_oa | False |
| best_oa_location.source.issn_l | 1552-5260 |
| best_oa_location.source.is_core | True |
| best_oa_location.source.is_in_doaj | False |
| best_oa_location.source.display_name | Alzheimer s & Dementia |
| best_oa_location.source.host_organization | https://openalex.org/P4310320595 |
| best_oa_location.source.host_organization_name | Wiley |
| best_oa_location.source.host_organization_lineage | https://openalex.org/P4310320595 |
| best_oa_location.license | cc-by |
| best_oa_location.pdf_url | |
| best_oa_location.version | publishedVersion |
| best_oa_location.raw_type | journal-article |
| best_oa_location.license_id | https://openalex.org/licenses/cc-by |
| best_oa_location.is_accepted | True |
| best_oa_location.is_published | True |
| best_oa_location.raw_source_name | Alzheimer's & Dementia |
| best_oa_location.landing_page_url | https://doi.org/10.1002/alz.092254 |
| primary_location.id | doi:10.1002/alz.092254 |
| primary_location.is_oa | True |
| primary_location.source.id | https://openalex.org/S108427512 |
| primary_location.source.issn | 1552-5260, 1552-5279 |
| primary_location.source.type | journal |
| primary_location.source.is_oa | False |
| primary_location.source.issn_l | 1552-5260 |
| primary_location.source.is_core | True |
| primary_location.source.is_in_doaj | False |
| primary_location.source.display_name | Alzheimer s & Dementia |
| primary_location.source.host_organization | https://openalex.org/P4310320595 |
| primary_location.source.host_organization_name | Wiley |
| primary_location.source.host_organization_lineage | https://openalex.org/P4310320595 |
| primary_location.license | cc-by |
| primary_location.pdf_url | |
| primary_location.version | publishedVersion |
| primary_location.raw_type | journal-article |
| primary_location.license_id | https://openalex.org/licenses/cc-by |
| primary_location.is_accepted | True |
| primary_location.is_published | True |
| primary_location.raw_source_name | Alzheimer's & Dementia |
| primary_location.landing_page_url | https://doi.org/10.1002/alz.092254 |
| publication_date | 2024-12-01 |
| publication_year | 2024 |
| referenced_works_count | 0 |
| abstract_inverted_index.3 | 123 |
| abstract_inverted_index.a | 6, 10, 58, 253, 258, 271 |
| abstract_inverted_index.AD | 41, 70, 170, 242 |
| abstract_inverted_index.We | 163 |
| abstract_inverted_index.as | 173, 216, 257 |
| abstract_inverted_index.be | 316, 340 |
| abstract_inverted_index.by | 54, 97, 309 |
| abstract_inverted_index.in | 19, 114, 149, 197, 243, 252, 279, 342 |
| abstract_inverted_index.is | 5 |
| abstract_inverted_index.of | 16, 274, 289 |
| abstract_inverted_index.on | 34, 57, 74, 100, 249, 292 |
| abstract_inverted_index.or | 82, 109, 241 |
| abstract_inverted_index.to | 43, 50, 237, 270, 301, 318, 325 |
| abstract_inverted_index.we | 48, 285, 299 |
| abstract_inverted_index.721 | 75 |
| abstract_inverted_index.AD. | 280 |
| abstract_inverted_index.CA1 | 150 |
| abstract_inverted_index.CAA | 222 |
| abstract_inverted_index.DNA | 77 |
| abstract_inverted_index.For | 331 |
| abstract_inverted_index.PCs | 124 |
| abstract_inverted_index.The | 27 |
| abstract_inverted_index.aim | 49 |
| abstract_inverted_index.and | 38, 68, 120, 145, 151, 159, 178, 191, 208, 221, 264, 321 |
| abstract_inverted_index.are | 15, 132 |
| abstract_inverted_index.but | 131, 226 |
| abstract_inverted_index.can | 268 |
| abstract_inverted_index.for | 117, 125, 168, 183, 206, 306 |
| abstract_inverted_index.not | 133, 233 |
| abstract_inverted_index.our | 282 |
| abstract_inverted_index.tau | 174 |
| abstract_inverted_index.the | 21, 121, 198, 238, 275, 287, 327 |
| abstract_inverted_index.to: | 135 |
| abstract_inverted_index.was | 72, 78, 89, 112 |
| abstract_inverted_index.(AD) | 4 |
| abstract_inverted_index.1000 | 101 |
| abstract_inverted_index.APOE | 224 |
| abstract_inverted_index.CA1) | 199 |
| abstract_inverted_index.CAA. | 179 |
| abstract_inverted_index.From | 281 |
| abstract_inverted_index.GWAS | 56, 293 |
| abstract_inverted_index.age, | 118 |
| abstract_inverted_index.also | 227 |
| abstract_inverted_index.been | 234 |
| abstract_inverted_index.data | 65, 88 |
| abstract_inverted_index.from | 80 |
| abstract_inverted_index.gene | 319 |
| abstract_inverted_index.have | 232 |
| abstract_inverted_index.hits | 167, 182 |
| abstract_inverted_index.like | 186 |
| abstract_inverted_index.loci | 211, 230, 266, 314 |
| abstract_inverted_index.more | 332 |
| abstract_inverted_index.pTDP | 147, 207 |
| abstract_inverted_index.rely | 33 |
| abstract_inverted_index.risk | 329 |
| abstract_inverted_index.such | 172, 215 |
| abstract_inverted_index.that | 23, 231 |
| abstract_inverted_index.this | 25 |
| abstract_inverted_index.were | 200, 204 |
| abstract_inverted_index.will | 315, 339 |
| abstract_inverted_index.with | 9, 223 |
| abstract_inverted_index.(CAA) | 143 |
| abstract_inverted_index.Here, | 47 |
| abstract_inverted_index.PLINK | 115 |
| abstract_inverted_index.SNPs, | 225 |
| abstract_inverted_index.actin | 195 |
| abstract_inverted_index.based | 99 |
| abstract_inverted_index.every | 126 |
| abstract_inverted_index.first | 122 |
| abstract_inverted_index.panel | 105 |
| abstract_inverted_index.risk. | 46 |
| abstract_inverted_index.size, | 298 |
| abstract_inverted_index.sizes | 37 |
| abstract_inverted_index.small | 296 |
| abstract_inverted_index.type, | 146 |
| abstract_inverted_index.using | 91, 106 |
| abstract_inverted_index.value | 18 |
| abstract_inverted_index.which | 267 |
| abstract_inverted_index.(GWAS) | 32 |
| abstract_inverted_index.ARTAG. | 209 |
| abstract_inverted_index.Hirano | 160, 192 |
| abstract_inverted_index.Linear | 108 |
| abstract_inverted_index.Method | 63 |
| abstract_inverted_index.Result | 162 |
| abstract_inverted_index.bodies | 193 |
| abstract_inverted_index.causal | 328 |
| abstract_inverted_index.deeper | 272 |
| abstract_inverted_index.deeply | 59 |
| abstract_inverted_index.detect | 302 |
| abstract_inverted_index.found. | 201 |
| abstract_inverted_index.frozen | 81 |
| abstract_inverted_index.gender | 119 |
| abstract_inverted_index.genes. | 330 |
| abstract_inverted_index.gyrus, | 153 |
| abstract_inverted_index.linked | 236 |
| abstract_inverted_index.power. | 294 |
| abstract_inverted_index.sample | 36, 297 |
| abstract_inverted_index.stage, | 139 |
| abstract_inverted_index.strong | 11, 217 |
| abstract_inverted_index.(ARTAG) | 158 |
| abstract_inverted_index.(lcWGS) | 95 |
| abstract_inverted_index.Amyloid | 136 |
| abstract_inverted_index.Despite | 295 |
| abstract_inverted_index.Genetic | 13 |
| abstract_inverted_index.Genomes | 102 |
| abstract_inverted_index.Project | 103 |
| abstract_inverted_index.amyloid | 141, 176, 220 |
| abstract_inverted_index.between | 219 |
| abstract_inverted_index.bodies. | 161 |
| abstract_inverted_index.cohort. | 62 |
| abstract_inverted_index.dentate | 152 |
| abstract_inverted_index.disease | 3, 8 |
| abstract_inverted_index.genetic | 45, 87 |
| abstract_inverted_index.include | 130 |
| abstract_inverted_index.lesions | 71, 171, 185, 251, 277, 308 |
| abstract_inverted_index.limited | 134 |
| abstract_inverted_index.managed | 300 |
| abstract_inverted_index.observe | 286 |
| abstract_inverted_index.phases, | 137 |
| abstract_inverted_index.plaques | 177 |
| abstract_inverted_index.results | 284 |
| abstract_inverted_index.several | 229 |
| abstract_inverted_index.stages, | 188 |
| abstract_inverted_index.studies | 14, 31 |
| abstract_inverted_index.tissue. | 85 |
| abstract_inverted_index.underly | 24 |
| abstract_inverted_index.utilize | 39 |
| abstract_inverted_index.Abstract | 0 |
| abstract_inverted_index.GLIMPSE. | 107 |
| abstract_inverted_index.accurate | 290 |
| abstract_inverted_index.analysis | 324, 338 |
| abstract_inverted_index.benefits | 288 |
| abstract_inverted_index.cerebral | 140 |
| abstract_inverted_index.clinical | 40 |
| abstract_inverted_index.cohorts. | 345 |
| abstract_inverted_index.commonly | 333 |
| abstract_inverted_index.disease. | 26 |
| abstract_inverted_index.findings | 214 |
| abstract_inverted_index.followed | 96 |
| abstract_inverted_index.general. | 244 |
| abstract_inverted_index.hallmark | 67, 335 |
| abstract_inverted_index.included | 212, 228 |
| abstract_inverted_index.lesions, | 336 |
| abstract_inverted_index.logistic | 110 |
| abstract_inverted_index.observed | 278 |
| abstract_inverted_index.presence | 148 |
| abstract_inverted_index.publicly | 343 |
| abstract_inverted_index.rendered | 261 |
| abstract_inverted_index.severity | 144 |
| abstract_inverted_index.specific | 239 |
| abstract_inverted_index.tangles, | 175 |
| abstract_inverted_index.available | 344 |
| abstract_inverted_index.classical | 28 |
| abstract_inverted_index.collected | 73 |
| abstract_inverted_index.determine | 326 |
| abstract_inverted_index.diagnosis | 42 |
| abstract_inverted_index.eliminate | 51 |
| abstract_inverted_index.extracted | 79 |
| abstract_inverted_index.generated | 90 |
| abstract_inverted_index.performed | 113, 341 |
| abstract_inverted_index.phenotype | 240 |
| abstract_inverted_index.reference | 104 |
| abstract_inverted_index.regarding | 66 |
| abstract_inverted_index.relations | 218 |
| abstract_inverted_index.screening | 260 |
| abstract_inverted_index.subjected | 317 |
| abstract_inverted_index.Associated | 210, 313 |
| abstract_inverted_index.Background | 1 |
| abstract_inverted_index.Conclusion | 245 |
| abstract_inverted_index.Phenotypic | 64 |
| abstract_inverted_index.Suggestive | 202 |
| abstract_inverted_index.aggregates | 196 |
| abstract_inverted_index.angiopathy | 142 |
| abstract_inverted_index.cerebellar | 84 |
| abstract_inverted_index.contribute | 269 |
| abstract_inverted_index.correcting | 116 |
| abstract_inverted_index.discovered | 164, 205 |
| abstract_inverted_index.functional | 322 |
| abstract_inverted_index.imputation | 98 |
| abstract_inverted_index.individual | 276 |
| abstract_inverted_index.mechanisms | 22 |
| abstract_inverted_index.performing | 55 |
| abstract_inverted_index.phenotype. | 127 |
| abstract_inverted_index.phenotyped | 60 |
| abstract_inverted_index.phenotypes | 129 |
| abstract_inverted_index.phenotypic | 52, 247, 311 |
| abstract_inverted_index.previously | 235 |
| abstract_inverted_index.regression | 111 |
| abstract_inverted_index.sequencing | 94 |
| abstract_inverted_index.suggestive | 265 |
| abstract_inverted_index.validation | 323 |
| abstract_inverted_index.Braak‐NFT | 138 |
| abstract_inverted_index.association | 30 |
| abstract_inverted_index.co‐morbid | 69, 184 |
| abstract_inverted_index.eliminating | 310 |
| abstract_inverted_index.elucidating | 20 |
| abstract_inverted_index.information | 248 |
| abstract_inverted_index.investigate | 44 |
| abstract_inverted_index.phenotyping | 291 |
| abstract_inverted_index.preliminary | 259, 283 |
| abstract_inverted_index.replication | 337 |
| abstract_inverted_index.significant | 166, 181, 263, 304 |
| abstract_inverted_index.AD‐related | 307 |
| abstract_inverted_index.Furthermore, | 180 |
| abstract_inverted_index.Implementing | 246 |
| abstract_inverted_index.Investigated | 128 |
| abstract_inverted_index.associations | 203, 305 |
| abstract_inverted_index.degeneration | 190 |
| abstract_inverted_index.heterogenous | 7 |
| abstract_inverted_index.individuals. | 76 |
| abstract_inverted_index.investigated | 334 |
| abstract_inverted_index.Alzheimer’s | 2 |
| abstract_inverted_index.Subsequently, | 86 |
| abstract_inverted_index.age‐related | 156 |
| abstract_inverted_index.genome‐wide | 29, 165, 255, 262, 303 |
| abstract_inverted_index.heritability. | 12 |
| abstract_inverted_index.heterogeneity | 53 |
| abstract_inverted_index.investigation | 256 |
| abstract_inverted_index.irreplaceable | 17 |
| abstract_inverted_index.understanding | 273 |
| abstract_inverted_index.(extracellular | 194 |
| abstract_inverted_index.heterogeneity. | 312 |
| abstract_inverted_index.low‐coverage | 92 |
| abstract_inverted_index.prioritization | 320 |
| abstract_inverted_index.whole‐genome | 93 |
| abstract_inverted_index.granulovacuolar | 189 |
| abstract_inverted_index.paraffin‐fixed | 83 |
| abstract_inverted_index.spread/presence, | 155 |
| abstract_inverted_index.alpha‐synuclein | 154, 187 |
| abstract_inverted_index.ever‐increasing | 35 |
| abstract_inverted_index.hypothesis‐free | 254 |
| abstract_inverted_index.neuropathological | 61, 250 |
| abstract_inverted_index.well‐established | 169 |
| abstract_inverted_index.proof‐of‐concept | 213 |
| abstract_inverted_index.tau‐astrogliopathy | 157 |
| cited_by_percentile_year | |
| countries_distinct_count | 2 |
| institutions_distinct_count | 12 |
| citation_normalized_percentile.value | 0.40443795 |
| citation_normalized_percentile.is_in_top_1_percent | False |
| citation_normalized_percentile.is_in_top_10_percent | False |