Mannose receptor (MRC1) mediates uptake of dextran in macrophages via receptor-mediated endocytosis Article Swipe
 YOU?
·
· 2024
· Open Access
·
· DOI: https://doi.org/10.1101/2024.08.13.607841
YOU?
·
· 2024
· Open Access
·
· DOI: https://doi.org/10.1101/2024.08.13.607841
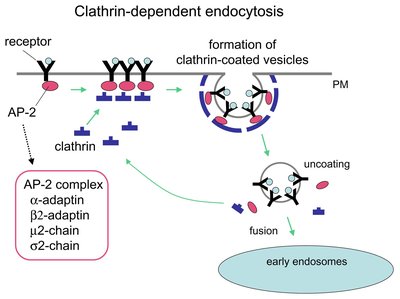
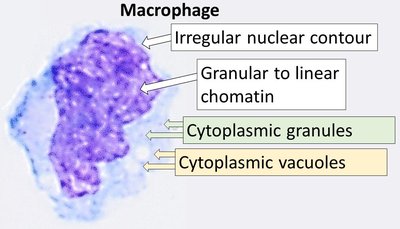
Macrophages maintain surveillance of their environment using receptor-mediated endocytosis and pinocytosis. Receptor-mediated endocytosis allows macrophages to recognize and internalize specific ligands whereas macropinocytosis non-selectively internalizes extracellular fluids and solutes. Here, CRISPR/Cas9 whole-genome screens were used to identify genes regulating constitutive and growth factor-stimulated dextran uptake in murine bone-marrow derived macrophages (BMDM). The endocytic mannose receptor c-type 1 ( Mrc1 , also known as CD206) was a top hit in the screen. Targeted gene disruptions of Mrc1 reduced dextran uptake but had little effect on uptake of Lucifer yellow, a fluid-phase marker. Other screen hits also differentially affected the uptake of dextran and Lucifer yellow, indicating the solutes are internalized by different mechanisms. We further deduced that BMDMs take up dextran via MRC1-mediated endocytosis by showing that competition with mannan, a ligand of MRC1, as well as treatment with Dyngo-4a, a dynamin inhibitor, reduced dextran uptake. Finally, we observed that IL4-treated BMDM internalize more dextran than untreated BMDM by upregulating MRC1 expression. These results demonstrate that dextran is not an effective marker for the bulk uptake of fluids and solutes by macropinocytosis since it is internalized by both macropinocytosis and receptor-mediated endocytosis in cells expressing MRC1. This report identifies numerous genes that regulate dextran internalization in primary murine macrophages and predicts cellular pathways and processes regulating MRC1. This work lays the groundwork for identifying specific genes and regulatory networks that regulate MRC1 expression and MRC1-mediated endocytosis in macrophages. Significance Statement Macrophages constantly survey and clear tissues by specifically and non-specifically internalizing debris and solutes. However, the molecular mechanisms and modes of regulation of these endocytic and macropinocytic processes are not well understood. Here, CRISPR/Cas9 whole genome screens were used to identify genes regulating uptake of dextran, a sugar polymer that is frequently used as a marker macropinocytosis, and compared with Lucifer yellow, a fluorescent dye with no known receptors. The authors identified the mannose receptor as well as other proteins regulating expression of the mannose receptor as top hits in the screen. Targeted disruption of Mrc1 , the gene that encodes mannose receptor, greatly diminished dextran uptake but had no effect on cellular uptake of Lucifer yellow. Furthermore, exposure to the cytokine IL4 upregulated mannose receptor expression on the cell surface and increased uptake of dextran with little effect on Lucifer yellow uptake. Studies seeking to understand regulation of macropinocytosis in macrophages will be confounded by the use of dextran as a fluid-phase marker. MRC1 is a marker of alternatively activated/anti-inflammatory macrophages and is a potential target for delivery of therapeutics to macrophages. This work provides the basis for mechanistic underpinning of how MRC1 contributes to the receptor-mediated uptake of carbohydrates and glycoproteins from the tissue milieu and distinguishes genes regulating receptor-mediated endocytosis from those regulating the bona fide fluid-phase uptake of fluids and solutes by macropinocytosis.
Related Topics
- Type
- preprint
- Language
- en
- Landing Page
- https://doi.org/10.1101/2024.08.13.607841
- https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdf
- OA Status
- green
- Cited By
- 1
- References
- 54
- Related Works
- 10
- OpenAlex ID
- https://openalex.org/W4401590302
Raw OpenAlex JSON
- OpenAlex ID
-
https://openalex.org/W4401590302Canonical identifier for this work in OpenAlex
- DOI
-
https://doi.org/10.1101/2024.08.13.607841Digital Object Identifier
- Title
-
Mannose receptor (MRC1) mediates uptake of dextran in macrophages via receptor-mediated endocytosisWork title
- Type
-
preprintOpenAlex work type
- Language
-
enPrimary language
- Publication year
-
2024Year of publication
- Publication date
-
2024-08-13Full publication date if available
- Authors
-
Jared Wollman, Kevin N. Wanniarachchi, Bijaya Pradhan, Lu Huang, Jason G. Kerkvliet, Adam D. Hoppe, Natalie W. ThiexList of authors in order
- Landing page
-
https://doi.org/10.1101/2024.08.13.607841Publisher landing page
- PDF URL
-
https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdfDirect link to full text PDF
- Open access
-
YesWhether a free full text is available
- OA status
-
greenOpen access status per OpenAlex
- OA URL
-
https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdfDirect OA link when available
- Concepts
-
Pinocytosis, Endocytosis, Mannose receptor, Cell biology, Biology, Lucifer yellow, Endocytic cycle, Receptor, Receptor-mediated endocytosis, Macrophage, Biochemistry, Intracellular, In vitro, Gap junctionTop concepts (fields/topics) attached by OpenAlex
- Cited by
-
1Total citation count in OpenAlex
- Citations by year (recent)
-
2025: 1Per-year citation counts (last 5 years)
- References (count)
-
54Number of works referenced by this work
- Related works (count)
-
10Other works algorithmically related by OpenAlex
Full payload
| id | https://openalex.org/W4401590302 |
|---|---|
| doi | https://doi.org/10.1101/2024.08.13.607841 |
| ids.doi | https://doi.org/10.1101/2024.08.13.607841 |
| ids.pmid | https://pubmed.ncbi.nlm.nih.gov/39211167 |
| ids.openalex | https://openalex.org/W4401590302 |
| fwci | 0.65076321 |
| type | preprint |
| title | Mannose receptor (MRC1) mediates uptake of dextran in macrophages via receptor-mediated endocytosis |
| biblio.issue | |
| biblio.volume | |
| biblio.last_page | |
| biblio.first_page | |
| topics[0].id | https://openalex.org/T10617 |
| topics[0].field.id | https://openalex.org/fields/13 |
| topics[0].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[0].score | 0.9984999895095825 |
| topics[0].domain.id | https://openalex.org/domains/1 |
| topics[0].domain.display_name | Life Sciences |
| topics[0].subfield.id | https://openalex.org/subfields/1307 |
| topics[0].subfield.display_name | Cell Biology |
| topics[0].display_name | Cellular transport and secretion |
| topics[1].id | https://openalex.org/T10993 |
| topics[1].field.id | https://openalex.org/fields/24 |
| topics[1].field.display_name | Immunology and Microbiology |
| topics[1].score | 0.9941999912261963 |
| topics[1].domain.id | https://openalex.org/domains/1 |
| topics[1].domain.display_name | Life Sciences |
| topics[1].subfield.id | https://openalex.org/subfields/2403 |
| topics[1].subfield.display_name | Immunology |
| topics[1].display_name | Complement system in diseases |
| topics[2].id | https://openalex.org/T10878 |
| topics[2].field.id | https://openalex.org/fields/13 |
| topics[2].field.display_name | Biochemistry, Genetics and Molecular Biology |
| topics[2].score | 0.9871000051498413 |
| topics[2].domain.id | https://openalex.org/domains/1 |
| topics[2].domain.display_name | Life Sciences |
| topics[2].subfield.id | https://openalex.org/subfields/1312 |
| topics[2].subfield.display_name | Molecular Biology |
| topics[2].display_name | CRISPR and Genetic Engineering |
| is_xpac | False |
| apc_list | |
| apc_paid | |
| concepts[0].id | https://openalex.org/C40017333 |
| concepts[0].level | 4 |
| concepts[0].score | 0.9284137487411499 |
| concepts[0].wikidata | https://www.wikidata.org/wiki/Q321362 |
| concepts[0].display_name | Pinocytosis |
| concepts[1].id | https://openalex.org/C28005876 |
| concepts[1].level | 3 |
| concepts[1].score | 0.8563479781150818 |
| concepts[1].wikidata | https://www.wikidata.org/wiki/Q189814 |
| concepts[1].display_name | Endocytosis |
| concepts[2].id | https://openalex.org/C2776205754 |
| concepts[2].level | 4 |
| concepts[2].score | 0.8195414543151855 |
| concepts[2].wikidata | https://www.wikidata.org/wiki/Q6750945 |
| concepts[2].display_name | Mannose receptor |
| concepts[3].id | https://openalex.org/C95444343 |
| concepts[3].level | 1 |
| concepts[3].score | 0.6622380018234253 |
| concepts[3].wikidata | https://www.wikidata.org/wiki/Q7141 |
| concepts[3].display_name | Cell biology |
| concepts[4].id | https://openalex.org/C86803240 |
| concepts[4].level | 0 |
| concepts[4].score | 0.5599753856658936 |
| concepts[4].wikidata | https://www.wikidata.org/wiki/Q420 |
| concepts[4].display_name | Biology |
| concepts[5].id | https://openalex.org/C2779486281 |
| concepts[5].level | 4 |
| concepts[5].score | 0.5251930952072144 |
| concepts[5].wikidata | https://www.wikidata.org/wiki/Q3163084 |
| concepts[5].display_name | Lucifer yellow |
| concepts[6].id | https://openalex.org/C79747257 |
| concepts[6].level | 4 |
| concepts[6].score | 0.46182897686958313 |
| concepts[6].wikidata | https://www.wikidata.org/wiki/Q189814 |
| concepts[6].display_name | Endocytic cycle |
| concepts[7].id | https://openalex.org/C170493617 |
| concepts[7].level | 2 |
| concepts[7].score | 0.4535329341888428 |
| concepts[7].wikidata | https://www.wikidata.org/wiki/Q208467 |
| concepts[7].display_name | Receptor |
| concepts[8].id | https://openalex.org/C117943116 |
| concepts[8].level | 4 |
| concepts[8].score | 0.4246430993080139 |
| concepts[8].wikidata | https://www.wikidata.org/wiki/Q1135604 |
| concepts[8].display_name | Receptor-mediated endocytosis |
| concepts[9].id | https://openalex.org/C2779244956 |
| concepts[9].level | 3 |
| concepts[9].score | 0.3672800660133362 |
| concepts[9].wikidata | https://www.wikidata.org/wiki/Q184204 |
| concepts[9].display_name | Macrophage |
| concepts[10].id | https://openalex.org/C55493867 |
| concepts[10].level | 1 |
| concepts[10].score | 0.26826226711273193 |
| concepts[10].wikidata | https://www.wikidata.org/wiki/Q7094 |
| concepts[10].display_name | Biochemistry |
| concepts[11].id | https://openalex.org/C79879829 |
| concepts[11].level | 2 |
| concepts[11].score | 0.20156419277191162 |
| concepts[11].wikidata | https://www.wikidata.org/wiki/Q5571762 |
| concepts[11].display_name | Intracellular |
| concepts[12].id | https://openalex.org/C202751555 |
| concepts[12].level | 2 |
| concepts[12].score | 0.06332707405090332 |
| concepts[12].wikidata | https://www.wikidata.org/wiki/Q221681 |
| concepts[12].display_name | In vitro |
| concepts[13].id | https://openalex.org/C158157758 |
| concepts[13].level | 3 |
| concepts[13].score | 0.0 |
| concepts[13].wikidata | https://www.wikidata.org/wiki/Q232438 |
| concepts[13].display_name | Gap junction |
| keywords[0].id | https://openalex.org/keywords/pinocytosis |
| keywords[0].score | 0.9284137487411499 |
| keywords[0].display_name | Pinocytosis |
| keywords[1].id | https://openalex.org/keywords/endocytosis |
| keywords[1].score | 0.8563479781150818 |
| keywords[1].display_name | Endocytosis |
| keywords[2].id | https://openalex.org/keywords/mannose-receptor |
| keywords[2].score | 0.8195414543151855 |
| keywords[2].display_name | Mannose receptor |
| keywords[3].id | https://openalex.org/keywords/cell-biology |
| keywords[3].score | 0.6622380018234253 |
| keywords[3].display_name | Cell biology |
| keywords[4].id | https://openalex.org/keywords/biology |
| keywords[4].score | 0.5599753856658936 |
| keywords[4].display_name | Biology |
| keywords[5].id | https://openalex.org/keywords/lucifer-yellow |
| keywords[5].score | 0.5251930952072144 |
| keywords[5].display_name | Lucifer yellow |
| keywords[6].id | https://openalex.org/keywords/endocytic-cycle |
| keywords[6].score | 0.46182897686958313 |
| keywords[6].display_name | Endocytic cycle |
| keywords[7].id | https://openalex.org/keywords/receptor |
| keywords[7].score | 0.4535329341888428 |
| keywords[7].display_name | Receptor |
| keywords[8].id | https://openalex.org/keywords/receptor-mediated-endocytosis |
| keywords[8].score | 0.4246430993080139 |
| keywords[8].display_name | Receptor-mediated endocytosis |
| keywords[9].id | https://openalex.org/keywords/macrophage |
| keywords[9].score | 0.3672800660133362 |
| keywords[9].display_name | Macrophage |
| keywords[10].id | https://openalex.org/keywords/biochemistry |
| keywords[10].score | 0.26826226711273193 |
| keywords[10].display_name | Biochemistry |
| keywords[11].id | https://openalex.org/keywords/intracellular |
| keywords[11].score | 0.20156419277191162 |
| keywords[11].display_name | Intracellular |
| keywords[12].id | https://openalex.org/keywords/in-vitro |
| keywords[12].score | 0.06332707405090332 |
| keywords[12].display_name | In vitro |
| language | en |
| locations[0].id | doi:10.1101/2024.08.13.607841 |
| locations[0].is_oa | True |
| locations[0].source.id | https://openalex.org/S4306402567 |
| locations[0].source.issn | |
| locations[0].source.type | repository |
| locations[0].source.is_oa | False |
| locations[0].source.issn_l | |
| locations[0].source.is_core | False |
| locations[0].source.is_in_doaj | False |
| locations[0].source.display_name | bioRxiv (Cold Spring Harbor Laboratory) |
| locations[0].source.host_organization | https://openalex.org/I2750212522 |
| locations[0].source.host_organization_name | Cold Spring Harbor Laboratory |
| locations[0].source.host_organization_lineage | https://openalex.org/I2750212522 |
| locations[0].license | |
| locations[0].pdf_url | https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdf |
| locations[0].version | acceptedVersion |
| locations[0].raw_type | posted-content |
| locations[0].license_id | |
| locations[0].is_accepted | True |
| locations[0].is_published | False |
| locations[0].raw_source_name | |
| locations[0].landing_page_url | https://doi.org/10.1101/2024.08.13.607841 |
| locations[1].id | pmid:39211167 |
| locations[1].is_oa | False |
| locations[1].source.id | https://openalex.org/S4306525036 |
| locations[1].source.issn | |
| locations[1].source.type | repository |
| locations[1].source.is_oa | False |
| locations[1].source.issn_l | |
| locations[1].source.is_core | False |
| locations[1].source.is_in_doaj | False |
| locations[1].source.display_name | PubMed |
| locations[1].source.host_organization | https://openalex.org/I1299303238 |
| locations[1].source.host_organization_name | National Institutes of Health |
| locations[1].source.host_organization_lineage | https://openalex.org/I1299303238 |
| locations[1].license | |
| locations[1].pdf_url | |
| locations[1].version | publishedVersion |
| locations[1].raw_type | |
| locations[1].license_id | |
| locations[1].is_accepted | True |
| locations[1].is_published | True |
| locations[1].raw_source_name | bioRxiv : the preprint server for biology |
| locations[1].landing_page_url | https://pubmed.ncbi.nlm.nih.gov/39211167 |
| locations[2].id | pmh:oai:pubmedcentral.nih.gov:11360935 |
| locations[2].is_oa | True |
| locations[2].source.id | https://openalex.org/S2764455111 |
| locations[2].source.issn | |
| locations[2].source.type | repository |
| locations[2].source.is_oa | False |
| locations[2].source.issn_l | |
| locations[2].source.is_core | False |
| locations[2].source.is_in_doaj | False |
| locations[2].source.display_name | PubMed Central |
| locations[2].source.host_organization | https://openalex.org/I1299303238 |
| locations[2].source.host_organization_name | National Institutes of Health |
| locations[2].source.host_organization_lineage | https://openalex.org/I1299303238 |
| locations[2].license | |
| locations[2].pdf_url | |
| locations[2].version | submittedVersion |
| locations[2].raw_type | Text |
| locations[2].license_id | |
| locations[2].is_accepted | False |
| locations[2].is_published | False |
| locations[2].raw_source_name | bioRxiv |
| locations[2].landing_page_url | https://www.ncbi.nlm.nih.gov/pmc/articles/11360935 |
| indexed_in | crossref, pubmed |
| authorships[0].author.id | https://openalex.org/A5004774949 |
| authorships[0].author.orcid | |
| authorships[0].author.display_name | Jared Wollman |
| authorships[0].countries | US |
| authorships[0].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[0].affiliations[0].raw_affiliation_string | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[0].institutions[0].id | https://openalex.org/I177156846 |
| authorships[0].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[0].institutions[0].type | education |
| authorships[0].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[0].institutions[0].country_code | US |
| authorships[0].institutions[0].display_name | South Dakota State University |
| authorships[0].author_position | first |
| authorships[0].raw_author_name | Jared Wollman |
| authorships[0].is_corresponding | False |
| authorships[0].raw_affiliation_strings | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[1].author.id | https://openalex.org/A5106519780 |
| authorships[1].author.orcid | https://orcid.org/0009-0008-9903-7802 |
| authorships[1].author.display_name | Kevin N. Wanniarachchi |
| authorships[1].countries | US |
| authorships[1].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[1].affiliations[0].raw_affiliation_string | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[1].institutions[0].id | https://openalex.org/I177156846 |
| authorships[1].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[1].institutions[0].type | education |
| authorships[1].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[1].institutions[0].country_code | US |
| authorships[1].institutions[0].display_name | South Dakota State University |
| authorships[1].author_position | middle |
| authorships[1].raw_author_name | Kevin Wanniarachchi |
| authorships[1].is_corresponding | False |
| authorships[1].raw_affiliation_strings | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[2].author.id | https://openalex.org/A5106507726 |
| authorships[2].author.orcid | |
| authorships[2].author.display_name | Bijaya Pradhan |
| authorships[2].countries | US |
| authorships[2].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[2].affiliations[0].raw_affiliation_string | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[2].institutions[0].id | https://openalex.org/I177156846 |
| authorships[2].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[2].institutions[0].type | education |
| authorships[2].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[2].institutions[0].country_code | US |
| authorships[2].institutions[0].display_name | South Dakota State University |
| authorships[2].author_position | middle |
| authorships[2].raw_author_name | Bijaya Pradhan |
| authorships[2].is_corresponding | False |
| authorships[2].raw_affiliation_strings | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[3].author.id | https://openalex.org/A5038062860 |
| authorships[3].author.orcid | https://orcid.org/0000-0002-7820-2177 |
| authorships[3].author.display_name | Lu Huang |
| authorships[3].countries | US |
| authorships[3].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[3].affiliations[0].raw_affiliation_string | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[3].institutions[0].id | https://openalex.org/I177156846 |
| authorships[3].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[3].institutions[0].type | education |
| authorships[3].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[3].institutions[0].country_code | US |
| authorships[3].institutions[0].display_name | South Dakota State University |
| authorships[3].author_position | middle |
| authorships[3].raw_author_name | Lu Huang |
| authorships[3].is_corresponding | False |
| authorships[3].raw_affiliation_strings | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[4].author.id | https://openalex.org/A5113550000 |
| authorships[4].author.orcid | |
| authorships[4].author.display_name | Jason G. Kerkvliet |
| authorships[4].countries | US |
| authorships[4].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[4].affiliations[0].raw_affiliation_string | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[4].affiliations[1].institution_ids | https://openalex.org/I177156846 |
| authorships[4].affiliations[1].raw_affiliation_string | Chemistry, Biochemistry and Physics Department, South Dakota State University, Brookings, SD, USA |
| authorships[4].institutions[0].id | https://openalex.org/I177156846 |
| authorships[4].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[4].institutions[0].type | education |
| authorships[4].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[4].institutions[0].country_code | US |
| authorships[4].institutions[0].display_name | South Dakota State University |
| authorships[4].author_position | middle |
| authorships[4].raw_author_name | Jason G Kerkvliet |
| authorships[4].is_corresponding | False |
| authorships[4].raw_affiliation_strings | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA, Chemistry, Biochemistry and Physics Department, South Dakota State University, Brookings, SD, USA |
| authorships[5].author.id | https://openalex.org/A5024851455 |
| authorships[5].author.orcid | https://orcid.org/0000-0003-4180-0840 |
| authorships[5].author.display_name | Adam D. Hoppe |
| authorships[5].countries | US |
| authorships[5].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[5].affiliations[0].raw_affiliation_string | Chemistry, Biochemistry and Physics Department, South Dakota State University, Brookings, SD, USA |
| authorships[5].institutions[0].id | https://openalex.org/I177156846 |
| authorships[5].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[5].institutions[0].type | education |
| authorships[5].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[5].institutions[0].country_code | US |
| authorships[5].institutions[0].display_name | South Dakota State University |
| authorships[5].author_position | middle |
| authorships[5].raw_author_name | Adam D Hoppe |
| authorships[5].is_corresponding | False |
| authorships[5].raw_affiliation_strings | Chemistry, Biochemistry and Physics Department, South Dakota State University, Brookings, SD, USA |
| authorships[6].author.id | https://openalex.org/A5038932659 |
| authorships[6].author.orcid | https://orcid.org/0000-0001-7028-094X |
| authorships[6].author.display_name | Natalie W. Thiex |
| authorships[6].countries | US |
| authorships[6].affiliations[0].institution_ids | https://openalex.org/I177156846 |
| authorships[6].affiliations[0].raw_affiliation_string | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| authorships[6].institutions[0].id | https://openalex.org/I177156846 |
| authorships[6].institutions[0].ror | https://ror.org/015jmes13 |
| authorships[6].institutions[0].type | education |
| authorships[6].institutions[0].lineage | https://openalex.org/I177156846 |
| authorships[6].institutions[0].country_code | US |
| authorships[6].institutions[0].display_name | South Dakota State University |
| authorships[6].author_position | last |
| authorships[6].raw_author_name | Natalie W Thiex |
| authorships[6].is_corresponding | True |
| authorships[6].raw_affiliation_strings | Biology and Microbiology Department, South Dakota State University, Brookings, SD, USA |
| has_content.pdf | True |
| has_content.grobid_xml | False |
| is_paratext | False |
| open_access.is_oa | True |
| open_access.oa_url | https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdf |
| open_access.oa_status | green |
| open_access.any_repository_has_fulltext | False |
| created_date | 2025-10-10T00:00:00 |
| display_name | Mannose receptor (MRC1) mediates uptake of dextran in macrophages via receptor-mediated endocytosis |
| has_fulltext | False |
| is_retracted | False |
| updated_date | 2025-11-06T06:51:31.235846 |
| primary_topic.id | https://openalex.org/T10617 |
| primary_topic.field.id | https://openalex.org/fields/13 |
| primary_topic.field.display_name | Biochemistry, Genetics and Molecular Biology |
| primary_topic.score | 0.9984999895095825 |
| primary_topic.domain.id | https://openalex.org/domains/1 |
| primary_topic.domain.display_name | Life Sciences |
| primary_topic.subfield.id | https://openalex.org/subfields/1307 |
| primary_topic.subfield.display_name | Cell Biology |
| primary_topic.display_name | Cellular transport and secretion |
| related_works | https://openalex.org/W1998284209, https://openalex.org/W2160431708, https://openalex.org/W3193270327, https://openalex.org/W4210371115, https://openalex.org/W2139436990, https://openalex.org/W3023999421, https://openalex.org/W2120834164, https://openalex.org/W2107801352, https://openalex.org/W4206212907, https://openalex.org/W2804243913 |
| cited_by_count | 1 |
| counts_by_year[0].year | 2025 |
| counts_by_year[0].cited_by_count | 1 |
| locations_count | 3 |
| best_oa_location.id | doi:10.1101/2024.08.13.607841 |
| best_oa_location.is_oa | True |
| best_oa_location.source.id | https://openalex.org/S4306402567 |
| best_oa_location.source.issn | |
| best_oa_location.source.type | repository |
| best_oa_location.source.is_oa | False |
| best_oa_location.source.issn_l | |
| best_oa_location.source.is_core | False |
| best_oa_location.source.is_in_doaj | False |
| best_oa_location.source.display_name | bioRxiv (Cold Spring Harbor Laboratory) |
| best_oa_location.source.host_organization | https://openalex.org/I2750212522 |
| best_oa_location.source.host_organization_name | Cold Spring Harbor Laboratory |
| best_oa_location.source.host_organization_lineage | https://openalex.org/I2750212522 |
| best_oa_location.license | |
| best_oa_location.pdf_url | https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdf |
| best_oa_location.version | acceptedVersion |
| best_oa_location.raw_type | posted-content |
| best_oa_location.license_id | |
| best_oa_location.is_accepted | True |
| best_oa_location.is_published | False |
| best_oa_location.raw_source_name | |
| best_oa_location.landing_page_url | https://doi.org/10.1101/2024.08.13.607841 |
| primary_location.id | doi:10.1101/2024.08.13.607841 |
| primary_location.is_oa | True |
| primary_location.source.id | https://openalex.org/S4306402567 |
| primary_location.source.issn | |
| primary_location.source.type | repository |
| primary_location.source.is_oa | False |
| primary_location.source.issn_l | |
| primary_location.source.is_core | False |
| primary_location.source.is_in_doaj | False |
| primary_location.source.display_name | bioRxiv (Cold Spring Harbor Laboratory) |
| primary_location.source.host_organization | https://openalex.org/I2750212522 |
| primary_location.source.host_organization_name | Cold Spring Harbor Laboratory |
| primary_location.source.host_organization_lineage | https://openalex.org/I2750212522 |
| primary_location.license | |
| primary_location.pdf_url | https://www.biorxiv.org/content/biorxiv/early/2024/08/13/2024.08.13.607841.full.pdf |
| primary_location.version | acceptedVersion |
| primary_location.raw_type | posted-content |
| primary_location.license_id | |
| primary_location.is_accepted | True |
| primary_location.is_published | False |
| primary_location.raw_source_name | |
| primary_location.landing_page_url | https://doi.org/10.1101/2024.08.13.607841 |
| publication_date | 2024-08-13 |
| publication_year | 2024 |
| referenced_works | https://openalex.org/W2104603012, https://openalex.org/W2072645067, https://openalex.org/W2007647318, https://openalex.org/W2112417177, https://openalex.org/W2930463924, https://openalex.org/W2971784092, https://openalex.org/W2023455126, https://openalex.org/W2154283358, https://openalex.org/W2286099491, https://openalex.org/W2064815984, https://openalex.org/W2320391773, https://openalex.org/W2252568502, https://openalex.org/W2994423700, https://openalex.org/W2045796965, https://openalex.org/W1984011934, https://openalex.org/W1987642183, https://openalex.org/W2095083502, https://openalex.org/W2898972432, https://openalex.org/W2036176224, https://openalex.org/W2774116239, https://openalex.org/W2045435533, https://openalex.org/W2601885162, https://openalex.org/W2109491705, https://openalex.org/W3045318893, https://openalex.org/W2030233651, https://openalex.org/W2239190724, https://openalex.org/W4226166227, https://openalex.org/W2160431708, https://openalex.org/W2154598603, https://openalex.org/W2100122648, https://openalex.org/W3046844007, https://openalex.org/W2139946699, https://openalex.org/W2097851646, https://openalex.org/W2091161919, https://openalex.org/W4313377050, https://openalex.org/W1549239087, https://openalex.org/W2532360980, https://openalex.org/W2937555910, https://openalex.org/W1992189978, https://openalex.org/W2101407387, https://openalex.org/W3188359536, https://openalex.org/W2035033919, https://openalex.org/W2105006553, https://openalex.org/W2766771070, https://openalex.org/W2111695773, https://openalex.org/W2167279371, https://openalex.org/W1973260880, https://openalex.org/W2040283218, https://openalex.org/W1586250986, https://openalex.org/W2115235864, https://openalex.org/W2167454162, https://openalex.org/W2093079184, https://openalex.org/W2046002438, https://openalex.org/W1423560049 |
| referenced_works_count | 54 |
| abstract_inverted_index.( | 58 |
| abstract_inverted_index., | 60, 336 |
| abstract_inverted_index.1 | 57 |
| abstract_inverted_index.a | 66, 89, 130, 140, 286, 294, 302, 401, 406, 414 |
| abstract_inverted_index.We | 113 |
| abstract_inverted_index.an | 169 |
| abstract_inverted_index.as | 63, 134, 136, 293, 315, 317, 326, 400 |
| abstract_inverted_index.be | 393 |
| abstract_inverted_index.by | 110, 124, 158, 180, 186, 246, 395, 465 |
| abstract_inverted_index.in | 46, 69, 192, 205, 236, 329, 390 |
| abstract_inverted_index.is | 167, 184, 290, 405, 413 |
| abstract_inverted_index.it | 183 |
| abstract_inverted_index.no | 306, 349 |
| abstract_inverted_index.of | 4, 75, 86, 100, 132, 176, 260, 262, 284, 322, 334, 354, 374, 388, 398, 408, 419, 431, 439, 461 |
| abstract_inverted_index.on | 84, 351, 367, 379 |
| abstract_inverted_index.to | 16, 36, 279, 359, 385, 421, 435 |
| abstract_inverted_index.up | 119 |
| abstract_inverted_index.we | 147 |
| abstract_inverted_index.IL4 | 362 |
| abstract_inverted_index.The | 52, 309 |
| abstract_inverted_index.and | 10, 18, 28, 41, 102, 178, 189, 209, 213, 226, 233, 243, 248, 252, 258, 265, 297, 371, 412, 441, 447, 463 |
| abstract_inverted_index.are | 108, 268 |
| abstract_inverted_index.but | 80, 347 |
| abstract_inverted_index.dye | 304 |
| abstract_inverted_index.for | 172, 222, 417, 428 |
| abstract_inverted_index.had | 81, 348 |
| abstract_inverted_index.hit | 68 |
| abstract_inverted_index.how | 432 |
| abstract_inverted_index.not | 168, 269 |
| abstract_inverted_index.the | 70, 98, 106, 173, 220, 255, 312, 323, 330, 337, 360, 368, 396, 426, 436, 444, 456 |
| abstract_inverted_index.top | 67, 327 |
| abstract_inverted_index.use | 397 |
| abstract_inverted_index.via | 121 |
| abstract_inverted_index.was | 65 |
| abstract_inverted_index.BMDM | 151, 157 |
| abstract_inverted_index.MRC1 | 160, 231, 404, 433 |
| abstract_inverted_index.Mrc1 | 59, 76, 335 |
| abstract_inverted_index.This | 196, 217, 423 |
| abstract_inverted_index.also | 61, 95 |
| abstract_inverted_index.bona | 457 |
| abstract_inverted_index.both | 187 |
| abstract_inverted_index.bulk | 174 |
| abstract_inverted_index.cell | 369 |
| abstract_inverted_index.fide | 458 |
| abstract_inverted_index.from | 443, 453 |
| abstract_inverted_index.gene | 73, 338 |
| abstract_inverted_index.hits | 94, 328 |
| abstract_inverted_index.lays | 219 |
| abstract_inverted_index.more | 153 |
| abstract_inverted_index.take | 118 |
| abstract_inverted_index.than | 155 |
| abstract_inverted_index.that | 116, 126, 149, 165, 201, 229, 289, 339 |
| abstract_inverted_index.used | 35, 278, 292 |
| abstract_inverted_index.well | 135, 270, 316 |
| abstract_inverted_index.were | 34, 277 |
| abstract_inverted_index.will | 392 |
| abstract_inverted_index.with | 128, 138, 299, 305, 376 |
| abstract_inverted_index.work | 218, 424 |
| abstract_inverted_index.BMDMs | 117 |
| abstract_inverted_index.Here, | 30, 272 |
| abstract_inverted_index.MRC1, | 133 |
| abstract_inverted_index.MRC1. | 195, 216 |
| abstract_inverted_index.Other | 92 |
| abstract_inverted_index.These | 162 |
| abstract_inverted_index.basis | 427 |
| abstract_inverted_index.cells | 193 |
| abstract_inverted_index.clear | 244 |
| abstract_inverted_index.genes | 38, 200, 225, 281, 449 |
| abstract_inverted_index.known | 62, 307 |
| abstract_inverted_index.modes | 259 |
| abstract_inverted_index.other | 318 |
| abstract_inverted_index.since | 182 |
| abstract_inverted_index.sugar | 287 |
| abstract_inverted_index.their | 5 |
| abstract_inverted_index.these | 263 |
| abstract_inverted_index.those | 454 |
| abstract_inverted_index.using | 7 |
| abstract_inverted_index.whole | 274 |
| abstract_inverted_index.CD206) | 64 |
| abstract_inverted_index.allows | 14 |
| abstract_inverted_index.c-type | 56 |
| abstract_inverted_index.debris | 251 |
| abstract_inverted_index.effect | 83, 350, 378 |
| abstract_inverted_index.fluids | 27, 177, 462 |
| abstract_inverted_index.genome | 275 |
| abstract_inverted_index.growth | 42 |
| abstract_inverted_index.ligand | 131 |
| abstract_inverted_index.little | 82, 377 |
| abstract_inverted_index.marker | 171, 295, 407 |
| abstract_inverted_index.milieu | 446 |
| abstract_inverted_index.murine | 47, 207 |
| abstract_inverted_index.report | 197 |
| abstract_inverted_index.screen | 93 |
| abstract_inverted_index.survey | 242 |
| abstract_inverted_index.target | 416 |
| abstract_inverted_index.tissue | 445 |
| abstract_inverted_index.uptake | 45, 79, 85, 99, 175, 283, 346, 353, 373, 438, 460 |
| abstract_inverted_index.yellow | 381 |
| abstract_inverted_index.(BMDM). | 51 |
| abstract_inverted_index.Lucifer | 87, 103, 300, 355, 380 |
| abstract_inverted_index.Studies | 383 |
| abstract_inverted_index.authors | 310 |
| abstract_inverted_index.deduced | 115 |
| abstract_inverted_index.derived | 49 |
| abstract_inverted_index.dextran | 44, 78, 101, 120, 144, 154, 166, 203, 345, 375, 399 |
| abstract_inverted_index.dynamin | 141 |
| abstract_inverted_index.encodes | 340 |
| abstract_inverted_index.further | 114 |
| abstract_inverted_index.greatly | 343 |
| abstract_inverted_index.ligands | 21 |
| abstract_inverted_index.mannan, | 129 |
| abstract_inverted_index.mannose | 54, 313, 324, 341, 364 |
| abstract_inverted_index.marker. | 91, 403 |
| abstract_inverted_index.polymer | 288 |
| abstract_inverted_index.primary | 206 |
| abstract_inverted_index.reduced | 77, 143 |
| abstract_inverted_index.results | 163 |
| abstract_inverted_index.screen. | 71, 331 |
| abstract_inverted_index.screens | 33, 276 |
| abstract_inverted_index.seeking | 384 |
| abstract_inverted_index.showing | 125 |
| abstract_inverted_index.solutes | 107, 179, 464 |
| abstract_inverted_index.surface | 370 |
| abstract_inverted_index.tissues | 245 |
| abstract_inverted_index.uptake. | 145, 382 |
| abstract_inverted_index.whereas | 22 |
| abstract_inverted_index.yellow, | 88, 104, 301 |
| abstract_inverted_index.yellow. | 356 |
| abstract_inverted_index.Abstract | 0 |
| abstract_inverted_index.Finally, | 146 |
| abstract_inverted_index.However, | 254 |
| abstract_inverted_index.Targeted | 72, 332 |
| abstract_inverted_index.affected | 97 |
| abstract_inverted_index.cellular | 211, 352 |
| abstract_inverted_index.compared | 298 |
| abstract_inverted_index.cytokine | 361 |
| abstract_inverted_index.delivery | 418 |
| abstract_inverted_index.dextran, | 285 |
| abstract_inverted_index.exposure | 358 |
| abstract_inverted_index.identify | 37, 280 |
| abstract_inverted_index.maintain | 2 |
| abstract_inverted_index.networks | 228 |
| abstract_inverted_index.numerous | 199 |
| abstract_inverted_index.observed | 148 |
| abstract_inverted_index.pathways | 212 |
| abstract_inverted_index.predicts | 210 |
| abstract_inverted_index.proteins | 319 |
| abstract_inverted_index.provides | 425 |
| abstract_inverted_index.receptor | 55, 314, 325, 365 |
| abstract_inverted_index.regulate | 202, 230 |
| abstract_inverted_index.solutes. | 29, 253 |
| abstract_inverted_index.specific | 20, 224 |
| abstract_inverted_index.Dyngo-4a, | 139 |
| abstract_inverted_index.Statement | 239 |
| abstract_inverted_index.different | 111 |
| abstract_inverted_index.effective | 170 |
| abstract_inverted_index.endocytic | 53, 264 |
| abstract_inverted_index.increased | 372 |
| abstract_inverted_index.molecular | 256 |
| abstract_inverted_index.potential | 415 |
| abstract_inverted_index.processes | 214, 267 |
| abstract_inverted_index.receptor, | 342 |
| abstract_inverted_index.recognize | 17 |
| abstract_inverted_index.treatment | 137 |
| abstract_inverted_index.untreated | 156 |
| abstract_inverted_index.confounded | 394 |
| abstract_inverted_index.constantly | 241 |
| abstract_inverted_index.diminished | 344 |
| abstract_inverted_index.disruption | 333 |
| abstract_inverted_index.expressing | 194 |
| abstract_inverted_index.expression | 232, 321, 366 |
| abstract_inverted_index.frequently | 291 |
| abstract_inverted_index.groundwork | 221 |
| abstract_inverted_index.identified | 311 |
| abstract_inverted_index.identifies | 198 |
| abstract_inverted_index.indicating | 105 |
| abstract_inverted_index.inhibitor, | 142 |
| abstract_inverted_index.mechanisms | 257 |
| abstract_inverted_index.receptors. | 308 |
| abstract_inverted_index.regulating | 39, 215, 282, 320, 450, 455 |
| abstract_inverted_index.regulation | 261, 387 |
| abstract_inverted_index.regulatory | 227 |
| abstract_inverted_index.understand | 386 |
| abstract_inverted_index.CRISPR/Cas9 | 31, 273 |
| abstract_inverted_index.IL4-treated | 150 |
| abstract_inverted_index.Macrophages | 1, 240 |
| abstract_inverted_index.bone-marrow | 48 |
| abstract_inverted_index.competition | 127 |
| abstract_inverted_index.contributes | 434 |
| abstract_inverted_index.demonstrate | 164 |
| abstract_inverted_index.disruptions | 74 |
| abstract_inverted_index.endocytosis | 9, 13, 123, 191, 235, 452 |
| abstract_inverted_index.environment | 6 |
| abstract_inverted_index.expression. | 161 |
| abstract_inverted_index.fluid-phase | 90, 402, 459 |
| abstract_inverted_index.fluorescent | 303 |
| abstract_inverted_index.identifying | 223 |
| abstract_inverted_index.internalize | 19, 152 |
| abstract_inverted_index.macrophages | 15, 50, 208, 391, 411 |
| abstract_inverted_index.mechanisms. | 112 |
| abstract_inverted_index.mechanistic | 429 |
| abstract_inverted_index.understood. | 271 |
| abstract_inverted_index.upregulated | 363 |
| abstract_inverted_index.Furthermore, | 357 |
| abstract_inverted_index.Significance | 238 |
| abstract_inverted_index.constitutive | 40 |
| abstract_inverted_index.internalized | 109, 185 |
| abstract_inverted_index.internalizes | 25 |
| abstract_inverted_index.macrophages. | 237, 422 |
| abstract_inverted_index.pinocytosis. | 11 |
| abstract_inverted_index.specifically | 247 |
| abstract_inverted_index.surveillance | 3 |
| abstract_inverted_index.therapeutics | 420 |
| abstract_inverted_index.underpinning | 430 |
| abstract_inverted_index.upregulating | 159 |
| abstract_inverted_index.whole-genome | 32 |
| abstract_inverted_index.MRC1-mediated | 122, 234 |
| abstract_inverted_index.alternatively | 409 |
| abstract_inverted_index.carbohydrates | 440 |
| abstract_inverted_index.distinguishes | 448 |
| abstract_inverted_index.extracellular | 26 |
| abstract_inverted_index.glycoproteins | 442 |
| abstract_inverted_index.internalizing | 250 |
| abstract_inverted_index.differentially | 96 |
| abstract_inverted_index.macropinocytic | 266 |
| abstract_inverted_index.internalization | 204 |
| abstract_inverted_index.non-selectively | 24 |
| abstract_inverted_index.macropinocytosis | 23, 181, 188, 389 |
| abstract_inverted_index.non-specifically | 249 |
| abstract_inverted_index.Receptor-mediated | 12 |
| abstract_inverted_index.factor-stimulated | 43 |
| abstract_inverted_index.macropinocytosis, | 296 |
| abstract_inverted_index.macropinocytosis. | 466 |
| abstract_inverted_index.receptor-mediated | 8, 190, 437, 451 |
| abstract_inverted_index.activated/anti-inflammatory | 410 |
| cited_by_percentile_year.max | 95 |
| cited_by_percentile_year.min | 91 |
| corresponding_author_ids | https://openalex.org/A5038932659 |
| countries_distinct_count | 1 |
| institutions_distinct_count | 7 |
| corresponding_institution_ids | https://openalex.org/I177156846 |
| sustainable_development_goals[0].id | https://metadata.un.org/sdg/15 |
| sustainable_development_goals[0].score | 0.4300000071525574 |
| sustainable_development_goals[0].display_name | Life in Land |
| citation_normalized_percentile.value | 0.62321971 |
| citation_normalized_percentile.is_in_top_1_percent | False |
| citation_normalized_percentile.is_in_top_10_percent | False |